Succinylome analysis reveals the involvement of lysine succinylation in metabolism in pathogenic Mycobacterium tuberculosis
- PMID: 25605462
- PMCID: PMC4390261
- DOI: 10.1074/mcp.M114.045922
Succinylome analysis reveals the involvement of lysine succinylation in metabolism in pathogenic Mycobacterium tuberculosis
Abstract
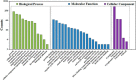
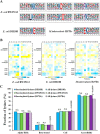
Mycobacterium tuberculosis (Mtb), the causative agent of human tuberculosis, remains one of the most prevalent human pathogens and a major cause of mortality worldwide. Metabolic network is a central mediator and defining feature of the pathogenicity of Mtb. Increasing evidence suggests that lysine succinylation dynamically regulates enzymes in carbon metabolism in both bacteria and human cells; however, its extent and function in Mtb remain unexplored. Here, we performed a global succinylome analysis of the virulent Mtb strain H37Rv by using high accuracy nano-LC-MS/MS in combination with the enrichment of succinylated peptides from digested cell lysates and subsequent peptide identification. In total, 1545 lysine succinylation sites on 626 proteins were identified in this pathogen. The identified succinylated proteins are involved in various biological processes and a large proportion of the succinylation sites are present on proteins in the central metabolism pathway. Site-specific mutations showed that succinylation is a negative regulatory modification on the enzymatic activity of acetyl-CoA synthetase. Molecular dynamics simulations demonstrated that succinylation affects the conformational stability of acetyl-CoA synthetase, which is critical for its enzymatic activity. Further functional studies showed that CobB, a sirtuin-like deacetylase in Mtb, functions as a desuccinylase of acetyl-CoA synthetase in in vitro assays. Together, our findings reveal widespread roles for lysine succinylation in regulating metabolism and diverse processes in Mtb. Our data provide a rich resource for functional analyses of lysine succinylation and facilitate the dissection of metabolic networks in this life-threatening pathogen.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures






Similar articles
-
First succinyl-proteome profiling of extensively drug-resistant Mycobacterium tuberculosis revealed involvement of succinylation in cellular physiology.J Proteome Res. 2015 Jan 2;14(1):107-19. doi: 10.1021/pr500859a. Epub 2014 Nov 12. J Proteome Res. 2015. PMID: 25363132
-
Lysine Succinylation Contributes to Aflatoxin Production and Pathogenicity in Aspergillus flavus.Mol Cell Proteomics. 2018 Mar;17(3):457-471. doi: 10.1074/mcp.RA117.000393. Epub 2018 Jan 3. Mol Cell Proteomics. 2018. PMID: 29298838 Free PMC article.
-
Global Proteome Analyses of Lysine Acetylation and Succinylation Reveal the Widespread Involvement of both Modification in Metabolism in the Embryo of Germinating Rice Seed.J Proteome Res. 2016 Mar 4;15(3):879-90. doi: 10.1021/acs.jproteome.5b00805. Epub 2016 Jan 28. J Proteome Res. 2016. PMID: 26767346
-
Histone succinylation and its function on the nucleosome.J Cell Mol Med. 2021 Aug;25(15):7101-7109. doi: 10.1111/jcmm.16676. Epub 2021 Jun 23. J Cell Mol Med. 2021. PMID: 34160884 Free PMC article. Review.
-
The Mystery of Extramitochondrial Proteins Lysine Succinylation.Int J Mol Sci. 2021 Jun 4;22(11):6085. doi: 10.3390/ijms22116085. Int J Mol Sci. 2021. PMID: 34199982 Free PMC article. Review.
Cited by
-
Metabolic Functions of Lysine 2-Hydroxyisobutyrylation.Cureus. 2020 Aug 11;12(8):e9651. doi: 10.7759/cureus.9651. Cureus. 2020. PMID: 32923251 Free PMC article. Review.
-
Lysine Propionylation is a Widespread Post-Translational Modification Involved in Regulation of Photosynthesis and Metabolism in Cyanobacteria.Int J Mol Sci. 2019 Sep 26;20(19):4792. doi: 10.3390/ijms20194792. Int J Mol Sci. 2019. PMID: 31561603 Free PMC article.
-
Comprehensive analysis of the succinylome in Vero cells infected with peste des petits ruminants virus Nigeria 75/1 vaccine strain.BMC Vet Res. 2025 Jan 30;21(1):45. doi: 10.1186/s12917-025-04496-3. BMC Vet Res. 2025. PMID: 39885502 Free PMC article.
-
Recent Contributions of Proteomics to Our Understanding of Reversible Nε-Lysine Acylation in Bacteria.J Proteome Res. 2024 Aug 2;23(8):2733-2749. doi: 10.1021/acs.jproteome.3c00912. Epub 2024 Mar 5. J Proteome Res. 2024. PMID: 38442041 Free PMC article. Review.
-
A Comprehensive Analysis of the Lysine Acetylome in the Aquatic Animals Pathogenic Bacterium Vibrio mimicus.Front Microbiol. 2022 Feb 17;13:816968. doi: 10.3389/fmicb.2022.816968. eCollection 2022. Front Microbiol. 2022. PMID: 35250932 Free PMC article.
References
-
- Miao J., Lawrence M., Jeffers V., Zhao F., Parker D., Ge Y., Sullivan W. J., Jr., Cui L. (2013) Extensive lysine acetylation occurs in evolutionarily conserved metabolic pathways and parasite-specific functions during Plasmodium falciparum intraerythrocytic development. Mol. Microbiol. 89, 660–675 - PMC - PubMed
-
- Zhao S., Xu W., Jiang W., Yu W., Lin Y., Zhang T., Yao J., Zhou L., Zeng Y., Li H., Li Y., Shi J., An W., Hancock S. M., He F., Qin L., Chin J., Yang P., Chen X., Lei Q., Xiong Y., Guan K. L. (2010) Regulation of cellular metabolism by protein lysine acetylation. Science 327, 1000–1004 - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

