Identification of the long non-coding RNA POU3F3 in plasma as a novel biomarker for diagnosis of esophageal squamous cell carcinoma
- PMID: 25608466
- PMCID: PMC4631113
- DOI: 10.1186/1476-4598-14-3
Identification of the long non-coding RNA POU3F3 in plasma as a novel biomarker for diagnosis of esophageal squamous cell carcinoma
Abstract
Background: Recent studies have demonstrated that long non-coding RNAs (lncRNAs) were present in the blood of cancer patients and have shown great potential as powerful and non-invasive tumor markers. However, little is known about the value of lncRNAs in the diagnosis of esophageal squamous cell carcinoma (ESCC). We hypothesized that ESCC-related lncRNAs might be released into the circulation during tumor initiation and could be utilized to detect and monitor ESCC.
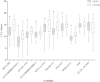
Methods: Ten lncRNAs (HOTAIR, AFAP1-AS1, POU3F3, HNF1A-AS1, 91H, PlncRNA1, SPRY4-IT1, ENST00000435885.1, XLOC_013104 and ENST00000547963.1) which previously found to be differently expressed in esophageal cancer were selected as candidate targets for subsequent circulating lncRNA assay. A four-stage exploratory study was conducted to test the hypothesis: (1) optimization of detected method to accurately and reproducibly measure ESCC-related lncRNAs in plasma and serum; (2) evaluation of the stability of circulating lncRNAs in human plasma or serum; (3) exploration the origin of ESCC-related lncRNAs in vitro and in vivo; (4) evaluation the diagnostic power of circulating lncRNAs for ESCC.
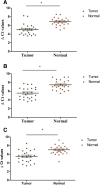
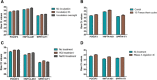
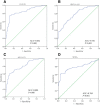
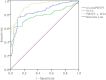
Results: ESCC-related lncRNAs were detectable and stable in plasma of cancer patients, and derived largely from ESCC tumor cells. Furthermore, plasma levels of POU3F3, HNF1A-AS1 and SPRY4-IT1 were significantly higher in ESCC patients compared with normal controls. By receiver operating characteristic curve (ROC) analysis, among the three lncRNAs investigated, plasma POU3F3 provided the highest diagnostic performance for detection of ESCC (the area under the ROC curve (AUC), 0.842; p < 0.001; sensitivity, 72.8%; specificity, 89.4%). Moreover, use of POU3F3 and SCCA in combination could provide a more effective diagnosis performance (AUC, 0.926, p < 0.001, sensitivity, 85.7%; specificity, 81.4%). Most importantly, this combination was effective to detect ESCC at an early stage (80.8%).
Conclusions: Plasma POU3F3 could serve as a potential biomarker for diagnosis of ESCC, and the combination of POU3F3 and SCCA was more efficient for ESCC detection, in particular for early tumor screening.
Figures


Similar articles
-
Three Circulating LncRNA Predict Early Progress of Esophageal Squamous Cell Carcinoma.Cell Physiol Biochem. 2016;40(1-2):117-125. doi: 10.1159/000452529. Epub 2016 Nov 18. Cell Physiol Biochem. 2016. PMID: 27855375
-
Serum HOTAIR as a novel diagnostic biomarker for esophageal squamous cell carcinoma.Mol Cancer. 2017 Apr 4;16(1):75. doi: 10.1186/s12943-017-0643-6. Mol Cancer. 2017. PMID: 28376832 Free PMC article.
-
Long noncoding RNA, tissue differentiation-inducing nonprotein coding RNA is upregulated and promotes development of esophageal squamous cell carcinoma.Dis Esophagus. 2016 Nov;29(8):950-958. doi: 10.1111/dote.12436. Epub 2016 Feb 2. Dis Esophagus. 2016. PMID: 26833746
-
The role of long non-coding RNA AFAP1-AS1 in human malignant tumors.Pathol Res Pract. 2018 Oct;214(10):1524-1531. doi: 10.1016/j.prp.2018.08.014. Epub 2018 Aug 20. Pathol Res Pract. 2018. PMID: 30173945 Review.
-
Noncoding RNA Expression Aberration Is Associated with Cancer Progression and Is a Potential Biomarker in Esophageal Squamous Cell Carcinoma.Int J Mol Sci. 2015 Nov 24;16(11):27824-34. doi: 10.3390/ijms161126060. Int J Mol Sci. 2015. PMID: 26610479 Free PMC article. Review.
Cited by
-
Non-Coding RNAs Regulate Placental Trophoblast Function and Participate in Recurrent Abortion.Front Pharmacol. 2021 Apr 22;12:646521. doi: 10.3389/fphar.2021.646521. eCollection 2021. Front Pharmacol. 2021. PMID: 33967782 Free PMC article. Review.
-
Serum lncRNAs (CCAT2, LINC01133, LINC00511) with Squamous Cell Carcinoma Antigen Panel as Novel Non-Invasive Biomarkers for Detection of Cervical Squamous Carcinoma.Cancer Manag Res. 2020 Oct 1;12:9495-9502. doi: 10.2147/CMAR.S259586. eCollection 2020. Cancer Manag Res. 2020. PMID: 33061636 Free PMC article.
-
Long non-coding RNA ZNF667-AS1 retards the development of esophageal squamous cell carcinoma via modulation of microRNA-1290-mediated PRUNE2.Transl Oncol. 2022 Jul;21:101371. doi: 10.1016/j.tranon.2022.101371. Epub 2022 Apr 30. Transl Oncol. 2022. PMID: 35504176 Free PMC article.
-
DNA methylation contributes to silencing the expression of linc00086 in gastric cancer.Oncol Lett. 2018 Aug;16(2):1931-1936. doi: 10.3892/ol.2018.8868. Epub 2018 Jun 1. Oncol Lett. 2018. PMID: 30008886 Free PMC article.
-
Clinical significance of SPRY4-IT1 in efficacy and survival prediction in breast cancer patients undergoing neoadjuvant chemotherapy.Histol Histopathol. 2020 Apr;35(4):361-370. doi: 10.14670/HH-18-175. Epub 2019 Oct 22. Histol Histopathol. 2020. PMID: 31638266
References
-
- van Hagen P, Hulshof MC, van Lanschot JJ, Steyerberg EW, van Berge Henegouwen MI, Wijnhoven BP, Richel DJ, Nieuwenhuijzen GA, Hospers GA, Bonenkamp JJ, Cuesta MA, Blaisse RJ, Busch OR, ten Kate FJ, Creemers GJ, Punt CJ, Plukker JT, Verheul HM, Spillenaar Bilgen EJ, van Dekken H, van der Sangen MJ, Rozema T, Biermann K, Beukema JC, Piet AH, van Rij CM, Reinders JG, Tilanus HW, van der Gaast A, CROSS Group Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med. 2012;366:2074–2084. doi: 10.1056/NEJMoa1112088. - DOI - PubMed
-
- Hirajima S, Komatsu S, Ichikawa D, Takeshita H, Konishi H, Shiozaki A, Morimura R, Tsujiura M, Nagata H, Kawaguchi T, Arita T, Kubota T, Fujiwara H, Okamoto K, Otsuji E. Clinical impact of circulating miR-18a in plasma of patients with oesophageal squamous cell carcinoma. Br J Cancer. 2013;108:1822–1829. doi: 10.1038/bjc.2013.148. - DOI - PMC - PubMed
-
- Kosugi S, Nishimaki T, Kanda T, Nakagawa S, Ohashi M, Hatakeyama K. Clinical significance of serum carcinoembryonic antigen, carbohydrate antigen 19–9, and squamous cell carcinoma antigen levels in esophageal cancer patients. World J Surg. 2004;28:680–685. doi: 10.1007/s00268-004-6865-y. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

