DNA motion induced by electrokinetic flow near an Au coated nanopore surface as voltage controlled gate
- PMID: 25611963
- PMCID: PMC4326562
- DOI: 10.1088/0957-4484/26/6/065502
DNA motion induced by electrokinetic flow near an Au coated nanopore surface as voltage controlled gate
Abstract
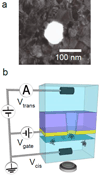
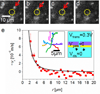
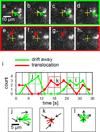
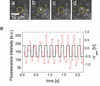
We used fluorescence microscopy to investigate the diffusion and drift motion of λ DNA molecules on an Au-coated membrane surface near nanopores, prior to their translocation through solid-state nanopores. With the capability of controlling electric potential at the Au surface as a gate voltage, Vgate, the motions of DNA molecules, which are presumably generated by electrokinetic flow, vary dramatically near the nanopores in our observations. We carefully investigate these DNA motions with different values of Vgate in order to alter the densities and polarities of the counterions, which are expected to change the flow speed or direction, respectively. Depending on Vgate, our observations have revealed the critical distance from a nanopore for DNA molecules to be attracted or repelled-DNA's anisotropic and unsteady drifting motions and accumulations of DNA molecules near the nanopore entrance. Further finite element method (FEM) numerical simulations indicate that the electrokinetic flow could qualitatively explain these unusual DNA motions near metal-collated gated nanopores. Finally, we demonstrate the possibility of controlling the speed and direction of DNA motion near or through a nanopore, as in the case of recapturing a single DNA molecule multiple times with alternating current voltages on the Vgate.
Figures


References
-
- Li J, Stein D, McMullan C, Branton D, Aziz MJ, Golovchenko JA. Ion-beam sculpting at nanometre length scales. Nature. 2001;412:166–169. - PubMed
-
- Storm AJ, Chen JH, Ling XS, Zandbergen HW, Dekker C. Fabrication of solid-state nanopores with single-nanometre precision. Nat. Mater. 2003;2:537–540. - PubMed
-
- Stein D, Li J, Golovchenko JA. Ion-beam sculpting time scales. Phys. Rev. Lett. 2002;89:276106. - PubMed
-
- Branton D, Deamer DW, Marziali A, Bayley H, Benner SA, Butler T, Di Ventra M, Garaj S, Hibbs A, Huang X, Jovanovich SB, Krstic PS, Lindsay S, Ling XS, Mastrangelo CH, Meller A, Oliver JS, Pershin YV, Ramsey JM, Riehn R, Soni GV, Tabard-Cossa V, Wanunu M, Wiggin M, Schloss JA. The potential and challenges of nanopore sequencing. Nat. Biotechnol. 2008;26:1146–1153. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
