High-resolution genetic mapping of allelic variants associated with cell wall chemistry in Populus
- PMID: 25613058
- PMCID: PMC4307895
- DOI: 10.1186/s12864-015-1215-z
High-resolution genetic mapping of allelic variants associated with cell wall chemistry in Populus
Abstract
Background: QTL cloning for the discovery of genes underlying polygenic traits has historically been cumbersome in long-lived perennial plants like Populus. Linkage disequilibrium-based association mapping has been proposed as a cloning tool, and recent advances in high-throughput genotyping and whole-genome resequencing enable marker saturation to levels sufficient for association mapping with no a priori candidate gene selection. Here, multiyear and multienvironment evaluation of cell wall phenotypes was conducted in an interspecific P. trichocarpa x P. deltoides pseudo-backcross mapping pedigree and two partially overlapping populations of unrelated P. trichocarpa genotypes using pyrolysis molecular beam mass spectrometry, saccharification, and/ or traditional wet chemistry. QTL mapping was conducted using a high-density genetic map with 3,568 SNP markers. As a fine-mapping approach, chromosome-wide association mapping targeting a QTL hot-spot on linkage group XIV was performed in the two P. trichocarpa populations. Both populations were genotyped using the 34 K Populus Infinium SNP array and whole-genome resequencing of one of the populations facilitated marker-saturation of candidate intervals for gene identification.
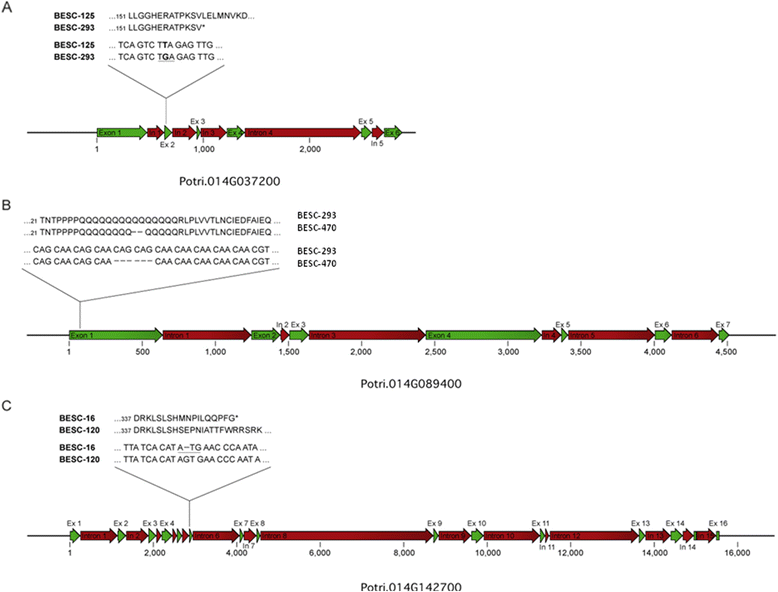
Results: Five QTLs ranging in size from 0.6 to 1.8 Mb were mapped on linkage group XIV for lignin content, syringyl to guaiacyl (S/G) ratio, 5- and 6-carbon sugars using the mapping pedigree. Six candidate loci exhibiting significant associations with phenotypes were identified within QTL intervals. These associations were reproducible across multiple environments, two independent genotyping platforms, and different plant growth stages. cDNA sequencing for allelic variants of three of the six loci identified polymorphisms leading to variable length poly glutamine (PolyQ) stretch in a transcription factor annotated as an ANGUSTIFOLIA C-terminus Binding Protein (CtBP) and premature stop codons in a KANADI transcription factor as well as a protein kinase. Results from protoplast transient expression assays suggested that each of the polymorphisms conferred allelic differences in the activation of cellulose, hemicelluloses, and lignin pathway marker genes.
Conclusion: This study illustrates the utility of complementary QTL and association mapping as tools for gene discovery with no a priori candidate gene selection. This proof of concept in a perennial organism opens up opportunities for discovery of novel genetic determinants of economically important but complex traits in plants.
Figures


Similar articles
-
Genetic architecture of growth traits in Populus revealed by integrated quantitative trait locus (QTL) analysis and association studies.New Phytol. 2016 Feb;209(3):1067-82. doi: 10.1111/nph.13695. Epub 2015 Oct 26. New Phytol. 2016. PMID: 26499329
-
Association genetics of traits controlling lignin and cellulose biosynthesis in black cottonwood (Populus trichocarpa, Salicaceae) secondary xylem.New Phytol. 2010 Oct;188(2):515-32. doi: 10.1111/j.1469-8137.2010.03415.x. Epub 2010 Sep 10. New Phytol. 2010. PMID: 20831625
-
Genome-wide association mapping for wood characteristics in Populus identifies an array of candidate single nucleotide polymorphisms.New Phytol. 2013 Nov;200(3):710-726. doi: 10.1111/nph.12422. Epub 2013 Jul 26. New Phytol. 2013. PMID: 23889164
-
QTL mapping using high-throughput sequencing.Methods Mol Biol. 2015;1284:257-85. doi: 10.1007/978-1-4939-2444-8_13. Methods Mol Biol. 2015. PMID: 25757777 Review.
-
Trait Mapping Approaches Through Association Analysis in Plants.Adv Biochem Eng Biotechnol. 2018;164:83-108. doi: 10.1007/10_2017_50. Adv Biochem Eng Biotechnol. 2018. PMID: 29511776 Review.
Cited by
-
Construction of a high-density genetic map and QTL mapping of leaf traits and plant growth in an interspecific F1 population of Catalpa bungei × Catalpa duclouxii Dode.BMC Plant Biol. 2019 Dec 30;19(1):596. doi: 10.1186/s12870-019-2207-y. BMC Plant Biol. 2019. PMID: 31888555 Free PMC article.
-
Joint linkage and association mapping of complex traits in shrub willow (Salix purpurea L.).Ann Bot. 2019 Oct 29;124(4):701-716. doi: 10.1093/aob/mcz047. Ann Bot. 2019. PMID: 31008500 Free PMC article.
-
Genetic variation of biomass recalcitrance in a natural Salix viminalis (L.) population.Biotechnol Biofuels. 2019 Jun 3;12:135. doi: 10.1186/s13068-019-1479-7. eCollection 2019. Biotechnol Biofuels. 2019. PMID: 31171936 Free PMC article.
-
A collection of genetically engineered Populus trees reveals wood biomass traits that predict glucose yield from enzymatic hydrolysis.Sci Rep. 2017 Nov 17;7(1):15798. doi: 10.1038/s41598-017-16013-0. Sci Rep. 2017. PMID: 29150693 Free PMC article.
-
Exon disruptive variants in Populus trichocarpa associated with wood properties exhibit distinct gene expression patterns.Plant Genome. 2025 Mar;18(1):e20541. doi: 10.1002/tpg2.20541. Epub 2024 Dec 4. Plant Genome. 2025. PMID: 39632472 Free PMC article.
References
-
- Dinus RJ, Payne P, Sewell MM, Chiang VL, Tuskan GA. Genetic modification of short rotation poplar wood: Properties for ethanol fuel and fiber production. Crit Rev Plant Sci. 2001;20:51–69. doi: 10.1016/S0735-2689(01)80012-5. - DOI
-
- Vermerris W, Saballos A, Ejeta G, Mosier NS, Ladisch MR. Molecular breeding to enhance ethanol production from corn and sorghum stover. Crop Sci. 2007;47:S142–S153. doi: 10.2135/cropsci2007.04.0013IPBS. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

