Memprot: a program to model the detergent corona around a membrane protein based on SEC-SAXS data
- PMID: 25615863
- PMCID: PMC4304689
- DOI: 10.1107/S1399004714016678
Memprot: a program to model the detergent corona around a membrane protein based on SEC-SAXS data
Abstract
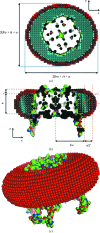
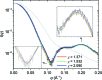
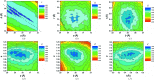
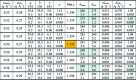
The application of small-angle X-ray scattering (SAXS) to structural investigations of transmembrane proteins in detergent solution has been hampered by two main inherent hurdles. On the one hand, the formation of a detergent corona around the hydrophobic region of the protein strongly modifies the scattering curve of the protein. On the other hand, free micelles of detergent without a precisely known concentration coexist with the protein-detergent complex in solution, therefore adding an uncontrolled signal. To gain robust structural information on such systems from SAXS data, in previous work, advantage was taken of the online combination of size-exclusion chromatography (SEC) and SAXS, and the detergent corona around aquaporin-0, a membrane protein of known structure, could be modelled. A precise geometrical model of the corona, shaped as an elliptical torus, was determined. Here, in order to better understand the correlations between the corona model parameters and to discuss the uniqueness of the model, this work was revisited by analyzing systematic SAXS simulations over a wide range of parameters of the torus.
Keywords: Memprot; SEC–SAXS; membrane proteins; small-angle X-ray scattering.
Figures



References
-
- Berthaud, A., Manzi, J., Pérez, J. & Mangenot, S. (2012). J. Am. Chem. Soc. 134, 10080–10088. - PubMed
-
- Breyton, C., Gabel, F., Lethier, M., Flayhan, A., Durand, G., Jault, J.-M., Juillan-Binard, C., Imbert, L., Moulin, M., Ravaud, S., Härtlein, M. & Ebel, C. (2013). Eur. Phys. J. E, 36, 71. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

