Cancer-associated protein kinase C mutations reveal kinase's role as tumor suppressor
- PMID: 25619690
- PMCID: PMC4313737
- DOI: 10.1016/j.cell.2015.01.001
Cancer-associated protein kinase C mutations reveal kinase's role as tumor suppressor
Abstract
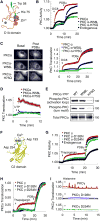
Protein kinase C (PKC) isozymes have remained elusive cancer targets despite the unambiguous tumor promoting function of their potent ligands, phorbol esters, and the prevalence of their mutations. We analyzed 8% of PKC mutations identified in human cancers and found that, surprisingly, most were loss of function and none were activating. Loss-of-function mutations occurred in all PKC subgroups and impeded second-messenger binding, phosphorylation, or catalysis. Correction of a loss-of-function PKCβ mutation by CRISPR-mediated genome editing in a patient-derived colon cancer cell line suppressed anchorage-independent growth and reduced tumor growth in a xenograft model. Hemizygous deletion promoted anchorage-independent growth, revealing that PKCβ is haploinsufficient for tumor suppression. Several mutations were dominant negative, suppressing global PKC signaling output, and bioinformatic analysis suggested that PKC mutations cooperate with co-occurring mutations in cancer drivers. These data establish that PKC isozymes generally function as tumor suppressors, indicating that therapies should focus on restoring, not inhibiting, PKC activity.
Copyright © 2015 Elsevier Inc. All rights reserved.
Figures
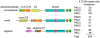



References
-
- Abbas T, White D, Hui L, Yoshida K, Foster DA, Bargonetti J. Inhibition of human p53 basal transcription by down-regulation of protein kinase Cdelta. J Biol Chem. 2004;279:9970–9977. - PubMed
-
- Alvaro V, Lévy L, Dubray C, Roche A, Peillon F, Quérat B, Joubert D. Invasive human pituitary tumors express a point-mutated alpha-protein kinase-C. J Clin Endocrinol Metab. 1993;77:1125–1129. - PubMed
-
- Barceló C, Paco N, Morell M, Alvarez-Moya B, Bota-Rabassedas N, Jaumot M, Vilardell F, Capella G, Agell N. Phosphorylation at Ser-181 of oncogenic KRAS is required for tumor growth. Cancer Res. 2014;74:1190–1199. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 12936/CRUK_/Cancer Research UK/United Kingdom
- R01 CA082683/CA/NCI NIH HHS/United States
- CA82683/CA/NCI NIH HHS/United States
- GM43154/GM/NIGMS NIH HHS/United States
- T32 GM007752/GM/NIGMS NIH HHS/United States
- NS080939/NS/NINDS NIH HHS/United States
- 15675/CRUK_/Cancer Research UK/United Kingdom
- P30 CA014195/CA/NCI NIH HHS/United States
- R37 GM043154/GM/NIGMS NIH HHS/United States
- R01 NS080939/NS/NINDS NIH HHS/United States
- R56 NS080939/NS/NINDS NIH HHS/United States
- R01 GM043154/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

