De novo assembly of transcriptome sequencing in Caragana korshinskii Kom. and characterization of EST-SSR markers
- PMID: 25629164
- PMCID: PMC4309406
- DOI: 10.1371/journal.pone.0115805
De novo assembly of transcriptome sequencing in Caragana korshinskii Kom. and characterization of EST-SSR markers
Abstract
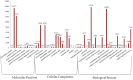
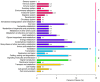
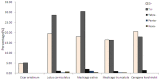
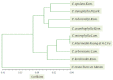
Caragana korshinskii Kom. is widely distributed in various habitats, including gravel desert, clay desert, fixed and semi-fixed sand, and saline land in the Asian and African deserts. To date, no previous genomic information or EST-SSR marker has been reported in Caragana Fabr. genus. In this study, more than two billion bases of high-quality sequence of C. korshinskii were generated by using illumina sequencing technology and demonstrated the de novo assembly and annotation of genes without prior genome information. These reads were assembled into 86,265 unigenes (mean length = 709 bp). The similarity search indicated that 33,955 and 21,978 unigenes showed significant similarities to known proteins from NCBI non-redundant and Swissprot protein databases, respectively. Among these annotated unigenes, 26,232 a unigenes were separately assigned to Gene Ontology (GO) database. When 22,756 unigenes searched against the Kyoto Encyclopedia of Genes and Genomes Pathway (KEGG) database, 5,598 unigenes were assigned to 5 main categories including 32 KEGG pathways. Among the main KEGG categories, metabolism was the biggest category (2,862, 43.7%), suggesting the active metabolic processes in the desert tree. In addition, a total of 19,150 EST-SSRs were identified from 15,484 unigenes, and the characterizations of EST-SSRs were further compared with other four species in Fabraceae. 126 potential marker sites were randomly selected to validate the assembly quality and develop EST-SSR markers. Among the 9 germplasms in Caranaga Fabr. genus, PCR success rate were 93.7% and the phylogenic tree was constructed based on the genotypic data. This research generated a substantial fraction of transcriptome sequences, which were very useful resources for gene annotation and discovery, molecular markers development, genome assembly and annotation. The EST-SSR markers identified and developed in this study will facilitate marker-assisted selection breeding.
Conflict of interest statement
Figures
Similar articles
-
De Novo Transcriptome Assembly of the Chinese Swamp Buffalo by RNA Sequencing and SSR Marker Discovery.PLoS One. 2016 Jan 14;11(1):e0147132. doi: 10.1371/journal.pone.0147132. eCollection 2016. PLoS One. 2016. PMID: 26766209 Free PMC article.
-
De novo assembly and characterization of bark transcriptome using Illumina sequencing and development of EST-SSR markers in rubber tree (Hevea brasiliensis Muell. Arg.).BMC Genomics. 2012 May 18;13:192. doi: 10.1186/1471-2164-13-192. BMC Genomics. 2012. PMID: 22607098 Free PMC article.
-
De novo assembly of the desert tree Haloxylon ammodendron (C. A. Mey.) based on RNA-Seq data provides insight into drought response, gene discovery and marker identification.BMC Genomics. 2014 Dec 15;15(1):1111. doi: 10.1186/1471-2164-15-1111. BMC Genomics. 2014. PMID: 25511667 Free PMC article.
-
Mining and Development of Novel SSR Markers Using Next Generation Sequencing (NGS) Data in Plants.Molecules. 2018 Feb 13;23(2):399. doi: 10.3390/molecules23020399. Molecules. 2018. PMID: 29438290 Free PMC article. Review.
-
A review of the major penaeid shrimp EST studies and the construction of a shrimp transcriptome database based on the ESTs from four penaeid shrimp.Mar Biotechnol (NY). 2011 Aug;13(4):608-21. doi: 10.1007/s10126-010-9286-y. Epub 2010 Apr 17. Mar Biotechnol (NY). 2011. PMID: 20401624 Review.
Cited by
-
The Potential Role of Genic-SSRs in Driving Ecological Adaptation Diversity in Caragana Plants.Int J Mol Sci. 2024 Feb 8;25(4):2084. doi: 10.3390/ijms25042084. Int J Mol Sci. 2024. PMID: 38396759 Free PMC article.
-
De Novo Transcriptome Assembly of the Chinese Swamp Buffalo by RNA Sequencing and SSR Marker Discovery.PLoS One. 2016 Jan 14;11(1):e0147132. doi: 10.1371/journal.pone.0147132. eCollection 2016. PLoS One. 2016. PMID: 26766209 Free PMC article.
-
Characterization of the Lycium barbarum fruit transcriptome and development of EST-SSR markers.PLoS One. 2017 Nov 10;12(11):e0187738. doi: 10.1371/journal.pone.0187738. eCollection 2017. PLoS One. 2017. PMID: 29125846 Free PMC article.
-
De Novo Sequencing and Transcriptome Analysis of Pleurotus eryngii subsp. tuoliensis (Bailinggu) Mycelia in Response to Cold Stimulation.Molecules. 2016 May 17;21(5):560. doi: 10.3390/molecules21050560. Molecules. 2016. PMID: 27196889 Free PMC article.
-
Characterizing the transcriptome and microsatellite markers for almond (Amygdalus communis L.) using the Illumina sequencing platform.Hereditas. 2017 Oct 19;155:14. doi: 10.1186/s41065-017-0049-x. eCollection 2018. Hereditas. 2017. PMID: 29075165 Free PMC article.
References
-
- Wang Z, Gao HW, Wu YQ, Han JG (2007) Genetic diversity and population structure of Caragana korshinskii revealed by AFLP. Crop Science 47: 1737–1743.
-
- Wang Z, Gao H (2008) Progress on genetic diversity of Genus Caragana germplasm resources. Journal of Plant Genetic Resources 9(3): 397–400.
-
- Yj Li, Zhao Z Sun Dx, Han G (2008) Hydrological physiological characteristics of Caragana korshinskii under water stress. Journal of Northwest Forestry University 23(3): 1–4.
-
- Yang DH, Song LY, Hu J, Yin WB, Li ZG, et al. (2012) Enhanced tolerance to NaCl and LiCl stresses by over-expressing Caragana korshinskii sodium/proton exchanger 1 (CkNHX1) and the hydrophilic C terminus is required for the activity of CkNHX1 in Atsos3–1 mutant and yeast. Biochem Biophys Res Commun 417: 732–737. 10.1016/j.bbrc.2011.12.023 - DOI - PubMed
Publication types
MeSH terms
Associated data
- Actions
- SRA/SRS724692
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

