Interaction of host cell microRNAs with the HCV RNA genome during infection of liver cells
- PMID: 25632937
- PMCID: PMC4832929
- DOI: 10.1055/s-0034-1397351
Interaction of host cell microRNAs with the HCV RNA genome during infection of liver cells
Abstract
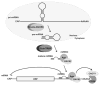
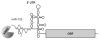
It has remained an enigma how hepatitis C viral (HCV) RNA can persist in the liver of infected patients for many decades. With the recent discovery of roles for microRNAs in gene expression, it was reported that the HCV RNA genome subverts liver-specific microRNA miR-122 to protect its 5' end from degradation by host cell exoribonucleases. Sequestration of miR-122 in cultured liver cells and in the liver of chimpanzees by small, modified antisense RNAs resulted in dramatic loss of HCV RNA and viral yield. This finding led to the first successful human trial in which subcutaneous administration of antisense molecules against miR-122 lowered viral yield in HCV patients, without the emergence of resistant virus. In this review, the authors summarize the molecular mechanism by which miR-122 protects the HCV RNA genome from degradation by exoribonucleases Xrn1 and Xrn2 and discuss the application of miR-122 antisense molecules in the clinic.
Thieme Medical Publishers 333 Seventh Avenue, New York, NY 10001, USA.
Figures

Similar articles
-
Dissecting the roles of the 5' exoribonucleases Xrn1 and Xrn2 in restricting hepatitis C virus replication.J Virol. 2015 May;89(9):4857-65. doi: 10.1128/JVI.03692-14. Epub 2015 Feb 11. J Virol. 2015. PMID: 25673723 Free PMC article.
-
Regulation of Hepatitis C Virus Genome Replication by Xrn1 and MicroRNA-122 Binding to Individual Sites in the 5' Untranslated Region.J Virol. 2015 Jun;89(12):6294-311. doi: 10.1128/JVI.03631-14. Epub 2015 Apr 8. J Virol. 2015. PMID: 25855736 Free PMC article.
-
Hepatitis C virus subverts liver-specific miR-122 to protect the viral genome from exoribonuclease Xrn2.Cell Host Microbe. 2014 Aug 13;16(2):257-264. doi: 10.1016/j.chom.2014.07.006. Cell Host Microbe. 2014. PMID: 25121753 Free PMC article.
-
Positive and negative modulation of viral and cellular mRNAs by liver-specific microRNA miR-122.Cold Spring Harb Symp Quant Biol. 2006;71:369-76. doi: 10.1101/sqb.2006.71.022. Cold Spring Harb Symp Quant Biol. 2006. PMID: 17381319 Review.
-
[Regulation of hepatitis C virus genome replication by microRNA-122].Uirusu. 2015;65(2):277-286. doi: 10.2222/jsv.65.277. Uirusu. 2015. PMID: 27760927 Review. Japanese.
Cited by
-
microRNA-99a Restricts Replication of Hepatitis C Virus by Targeting mTOR and de novo Lipogenesis.Viruses. 2020 Jun 27;12(7):696. doi: 10.3390/v12070696. Viruses. 2020. PMID: 32605105 Free PMC article.
-
Profiles of miRNAs in serum in severe acute drug induced liver injury and their prognostic significance.Liver Int. 2017 May;37(5):757-764. doi: 10.1111/liv.13312. Epub 2016 Nov 29. Liver Int. 2017. PMID: 27860186 Free PMC article.
-
Effects of oral implants with miR‑122‑modified cell sheets on rat bone marrow mesenchymal stem cells.Mol Med Rep. 2018 Jan;17(1):1537-1544. doi: 10.3892/mmr.2017.8094. Epub 2017 Nov 15. Mol Med Rep. 2018. PMID: 29257226 Free PMC article.
-
miR-122 does not impact recognition of the HCV genome by innate sensors of RNA but rather protects the 5' end from the cellular pyrophosphatases, DOM3Z and DUSP11.Nucleic Acids Res. 2018 Jun 1;46(10):5139-5158. doi: 10.1093/nar/gky273. Nucleic Acids Res. 2018. PMID: 29672716 Free PMC article.
-
LncRNAs in HCV Infection and HCV-Related Liver Disease.Int J Mol Sci. 2020 Mar 24;21(6):2255. doi: 10.3390/ijms21062255. Int J Mol Sci. 2020. PMID: 32214045 Free PMC article. Review.
References
-
- Hoofnagle JH. Course and outcome of hepatitis C. Hepatology. 2002;36(5, Suppl 1):S21–S29. - PubMed
-
- Lavanchy D. Evolving epidemiology of hepatitis C virus. Clin Microbiol Infect. 2011;17(2):107–115. - PubMed
-
- Casey LC, Lee WM. Hepatitis C virus therapy update 2013. Curr Opin Gastroenterol. 2013;29(3):243–249. - PubMed
-
- Bartenschlager R, Penin F, Lohmann V, André P. Assembly of infectious hepatitis C virus particles. Trends Microbiol. 2011;19(2):95–103. - PubMed
-
- Cannell IG, Kong YW, Bushell M. How do microRNAs regulate gene expression? Biochem Soc Trans. 2008;36(Pt 6):1224–1231. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

