Papillomaviruses: Viral evolution, cancer and evolutionary medicine
- PMID: 25634317
- PMCID: PMC4356112
- DOI: 10.1093/emph/eov003
Papillomaviruses: Viral evolution, cancer and evolutionary medicine
Abstract
Papillomaviruses (PVs) are a numerous family of small dsDNA viruses infecting virtually all mammals. PVs cause infections without triggering a strong immune response, and natural infection provides only limited protection against reinfection. Most PVs are part and parcel of the skin microbiota. In some cases, infections by certain PVs take diverse clinical presentations from highly productive self-limited warts to invasive cancers. We propose PVs as an excellent model system to study the evolutionary interactions between the immune system and pathogens causing chronic infections: genotypically, PVs are very diverse, with hundreds of different genotypes infecting skin and mucosa; phenotypically, they display extremely broad gradients and trade-offs between key phenotypic traits, namely productivity, immunogenicity, prevalence, oncogenicity and clinical presentation. Public health interventions have been launched to decrease the burden of PV-associated cancers, including massive vaccination against the most oncogenic human PVs, as well as systematic screening for PV chronic anogenital infections. Anti-PVs vaccines elicit protection against infection, induce cross-protection against closely related viruses and result in herd immunity. However, our knowledge on the ecological and intrapatient dynamics of PV infections remains fragmentary. We still need to understand how the novel anthropogenic selection pressures posed by vaccination and screening will affect viral circulation and epidemiology. We present here an overview of PV evolution and the connection between PV genotypes and the phenotypic, clinical manifestations of the diseases they cause. This differential link between viral evolution and the gradient cancer-warts-asymptomatic infections makes PVs a privileged playground for evolutionary medicine research.
Keywords: HPV; infection and cancer; molecular epidemiology; screening; vaccination; viral oncology; virus ecology cancer; virus evolution.
© The Author(s) 2015. Published by Oxford University Press on behalf of the Foundation for Evolution, Medicine, and Public Health.
Figures



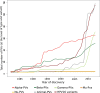

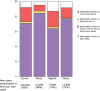
References
-
- Bravo IG, de Sanjose S, Gottschling M. The clinical importance of understanding the evolution of papillomaviruses. Trends Microbiol. 2010;18:432–8. - PubMed
-
- Garcia-Vallve S, Alonso A, Bravo IG. Papillomaviruses: different genes have different histories. Trends Microbiol. 2005;13:514–21. - PubMed
-
- Forman D, de Martel C, Lacey CJ, et al. Global burden of human papillomavirus and related diseases. Vaccine. 2012;30(Suppl 5):F12–23. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

