Global reorganization of the nuclear landscape in senescent cells
- PMID: 25640177
- PMCID: PMC4542308
- DOI: 10.1016/j.celrep.2014.12.055
Global reorganization of the nuclear landscape in senescent cells
Abstract
Cellular senescence has been implicated in tumor suppression, development, and aging and is accompanied by large-scale chromatin rearrangements, forming senescence-associated heterochromatic foci (SAHF). However, how the chromatin is reorganized during SAHF formation is poorly understood. Furthermore, heterochromatin formation in senescence appears to contrast with loss of heterochromatin in Hutchinson-Gilford progeria. We mapped architectural changes in genome organization in cellular senescence using Hi-C. Unexpectedly, we find a dramatic sequence- and lamin-dependent loss of local interactions in heterochromatin. This change in local connectivity resolves the paradox of opposing chromatin changes in senescence and progeria. In addition, we observe a senescence-specific spatial clustering of heterochromatic regions, suggesting a unique second step required for SAHF formation. Comparison of embryonic stem cells (ESCs), somatic cells, and senescent cells shows a unidirectional loss in local chromatin connectivity, suggesting that senescence is an endpoint of the continuous nuclear remodelling process during differentiation.
Copyright © 2015 The Authors. Published by Elsevier Inc. All rights reserved.
Figures

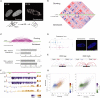

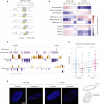
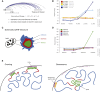
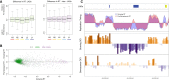

References
-
- Acosta J.C., O’Loghlen A., Banito A., Guijarro M.V., Augert A., Raguz S., Fumagalli M., Da Costa M., Brown C., Popov N. Chemokine signaling via the CXCR2 receptor reinforces senescence. Cell. 2008;133:1006–1018. - PubMed
-
- Bartkova J., Rezaei N., Liontos M., Karakaidos P., Kletsas D., Issaeva N., Vassiliou L.-V.F., Kolettas E., Niforou K., Zoumpourlis V.C. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature. 2006;444:633–637. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 095645/Z/11/Z/WT_/Wellcome Trust/United Kingdom
- 095645/WT_/Wellcome Trust/United Kingdom
- BB/K010867/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- G0801156/MRC_/Medical Research Council/United Kingdom
- BBS/E/B/000C0404/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources

