Molecular basis of the polyspecificity of P-glycoprotein (ABCB1): recent biochemical and structural studies
- PMID: 25640267
- PMCID: PMC7709800
- DOI: 10.1016/bs.acr.2014.10.003
Molecular basis of the polyspecificity of P-glycoprotein (ABCB1): recent biochemical and structural studies
Abstract
ABCB1 (P-glycoprotein/P-gp) is an ATP-binding cassette transporter well known for its association with multidrug resistance in cancer cells. Powered by the hydrolysis of ATP, it effluxes structurally diverse compounds. In this chapter, we discuss current views on the molecular basis of the substrate polyspecificity of P-gp. One of the features that accounts for this property is the structural flexibility observed in P-gp. Several X-ray crystal structures of mouse P-gp have been published recently in the absence of nucleotide, with and without bound inhibitors. All the structures are in an inward-facing conformation exhibiting different degrees of domain separation, thus revealing a highly flexible protein. Biochemical and biophysical studies also demonstrate this flexibility in mouse as well as human P-gp. Site-directed mutagenesis has revealed the existence of multiple transport-active binding sites in P-gp for a single substrate. Thus, drugs can bind at either primary or secondary sites. Biochemical, molecular modeling, and structure-activity relationship studies suggest a large, common drug-binding pocket with overlapping sites for different substrates. We propose that in addition to the structural flexibility, the molecular or chemical flexibility also contributes to the binding of substrates to multiple sites forming the basis of polyspecificity.
Keywords: ABC transporter; Drug-binding sites; Multidrug resistance; P-glycoprotein; Polyspecificity; Structural flexibility.
© 2015 Elsevier Inc. All rights reserved.
Figures
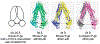



References
-
- Ambudkar SV, Cardarelli CO, Pashinsky I, & Stein WD (1997). Relation between the turnover number for vinblastine transport and for vinblastine-stimulated ATP hydrolysis by human P-glycoprotein. Journal of Biological Chemistry, 272, 21160–21166. - PubMed
-
- Ambudkar SV, Dey S, Hrycyna CA, Ramachandra M, Pastan I, & Gottesman MM (1999). Biochemical, cellular, and pharmacological aspects of the multidrug transporter. Annual Review of Pharmacology and Toxicology, 39, 361–398. - PubMed
-
- Ambudkar SV, Kim IW, & Sauna ZE (2006). The power of the pump: Mechanisms of action of P-glycoprotein (ABCB1). European Journal of Pharmaceutical Sciences, 27, 392–400. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

