Nanofibrous hydrogels with spatially patterned biochemical signals to control cell behavior
- PMID: 25640972
- PMCID: PMC4412590
- DOI: 10.1002/adma.201404993
Nanofibrous hydrogels with spatially patterned biochemical signals to control cell behavior
Abstract
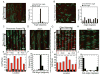
The ability to spatially pattern biochemical signals into nanofibrous materials using thiol-ene reactions of thiolated molecules to presented norbornene groups is demonstrated. This approach is used to pattern three molecules independently within one scaffold, to pattern molecules through the depth of a scaffold, and to spatially control cell adhesion and morphology.
Keywords: alignment; electrospinning; nanofibers; patterning; tissue engineering.
© 2015 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures



Similar articles
-
Hemocompatible surface of electrospun nanofibrous scaffolds by ATRP modification.Mater Sci Eng C Mater Biol Appl. 2013 Oct;33(7):3644-51. doi: 10.1016/j.msec.2013.04.048. Epub 2013 May 3. Mater Sci Eng C Mater Biol Appl. 2013. PMID: 23910260
-
Thiol-ene crosslinking polyamidoamine dendrimer-hyaluronic acid hydrogel system for biomedical applications.J Biomater Sci Polym Ed. 2016;27(8):743-57. doi: 10.1080/09205063.2016.1159473. Epub 2016 Mar 26. J Biomater Sci Polym Ed. 2016. PMID: 26923639
-
Development of nanofibrous scaffolds containing gum tragacanth/poly (ε-caprolactone) for application as skin scaffolds.Mater Sci Eng C Mater Biol Appl. 2015 Mar;48:71-9. doi: 10.1016/j.msec.2014.10.020. Epub 2014 Oct 13. Mater Sci Eng C Mater Biol Appl. 2015. PMID: 25579898
-
Design and manufacture of neural tissue engineering scaffolds using hyaluronic acid and polycaprolactone nanofibers with controlled porosity.Mater Sci Eng C Mater Biol Appl. 2016 Dec 1;69:380-7. doi: 10.1016/j.msec.2016.06.078. Epub 2016 Jun 25. Mater Sci Eng C Mater Biol Appl. 2016. PMID: 27612726
-
Electrospun hydrogels for dynamic culture systems: advantages, progress, and opportunities.Biomater Sci. 2021 Jun 21;9(12):4228-4245. doi: 10.1039/d0bm01588a. Epub 2021 Feb 1. Biomater Sci. 2021. PMID: 33522527 Free PMC article. Review.
Cited by
-
4D Biochemical Photocustomization of Hydrogel Scaffolds for Biomimetic Tissue Engineering.Acc Mater Res. 2023 Aug 25;4(8):704-715. doi: 10.1021/accountsmr.3c00062. Epub 2023 Jul 12. Acc Mater Res. 2023. PMID: 39071987 Free PMC article.
-
Hydration-Enhanced Lubricating Electrospun Nanofibrous Membranes Prevent Tissue Adhesion.Research (Wash D C). 2020 Mar 19;2020:4907185. doi: 10.34133/2020/4907185. eCollection 2020. Research (Wash D C). 2020. PMID: 32270140 Free PMC article.
-
Photo-tunable hydrogel mechanical heterogeneity informed by predictive transport kinetics model.Soft Matter. 2020 May 7;16(17):4131-4141. doi: 10.1039/d0sm00052c. Epub 2020 Mar 23. Soft Matter. 2020. PMID: 32202291 Free PMC article.
-
Progress in material design for biomedical applications.Proc Natl Acad Sci U S A. 2015 Nov 24;112(47):14444-51. doi: 10.1073/pnas.1516247112. Proc Natl Acad Sci U S A. 2015. PMID: 26598696 Free PMC article. Review.
-
Spatially and Temporally Controlled Hydrogels for Tissue Engineering.Mater Sci Eng R Rep. 2017 Sep;119:1-35. doi: 10.1016/j.mser.2017.07.001. Epub 2017 Jul 25. Mater Sci Eng R Rep. 2017. PMID: 29200661 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

