Mechanisms of gene duplication and amplification
- PMID: 25646380
- PMCID: PMC4315931
- DOI: 10.1101/cshperspect.a016592
Mechanisms of gene duplication and amplification
Abstract
Changes in gene copy number are among the most frequent mutational events in all genomes and were among the mutations for which a physical basis was first known. Yet mechanisms of gene duplication remain uncertain because formation rates are difficult to measure and mechanisms may vary with position in a genome. Duplications are compared here to deletions, which seem formally similar but can arise at very different rates by distinct mechanisms. Methods of assessing duplication rates and dependencies are described with several proposed formation mechanisms. Emphasis is placed on duplications formed in extensively studied experimental situations. Duplications studied in microbes are compared with those observed in metazoan cells, specifically those in genomes of cancer cells. Duplications, and especially their derived amplifications, are suggested to form by multistep processes often under positive selection for increased copy number.
Copyright © 2015 Cold Spring Harbor Laboratory Press; all rights reserved.
Figures
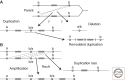
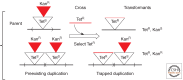

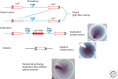
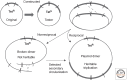


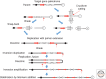
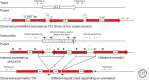
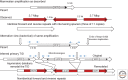
References
-
- Albertson DG 2006. Gene amplification in cancer. Trends Genet 22: 447–455. - PubMed
-
- Anderson RP, Roth JR 1977. Tandem genetic duplications in phage and bacteria. Annu Rev Microbiol 31: 473–505. - PubMed
-
- Anderson RP, Roth JR 1978. Tandem chromosomal duplications in Salmonella typhimurium: Fusion of histidine genes to novel promoters. J Mol Biol 119: 147–166. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
