The evolution of drug resistance in clinical isolates of Candida albicans
- PMID: 25646566
- PMCID: PMC4383195
- DOI: 10.7554/eLife.00662
The evolution of drug resistance in clinical isolates of Candida albicans
Abstract
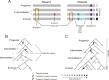
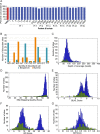
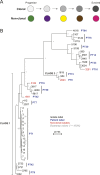
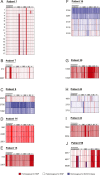
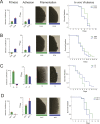
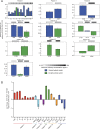
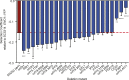
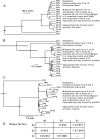
Candida albicans is both a member of the healthy human microbiome and a major pathogen in immunocompromised individuals. Infections are typically treated with azole inhibitors of ergosterol biosynthesis often leading to drug resistance. Studies in clinical isolates have implicated multiple mechanisms in resistance, but have focused on large-scale aberrations or candidate genes, and do not comprehensively chart the genetic basis of adaptation. Here, we leveraged next-generation sequencing to analyze 43 isolates from 11 oral candidiasis patients. We detected newly selected mutations, including single-nucleotide polymorphisms (SNPs), copy-number variations and loss-of-heterozygosity (LOH) events. LOH events were commonly associated with acquired resistance, and SNPs in 240 genes may be related to host adaptation. Conversely, most aneuploidies were transient and did not correlate with drug resistance. Our analysis also shows that isolates also varied in adherence, filamentation, and virulence. Our work reveals new molecular mechanisms underlying the evolution of drug resistance and host adaptation.
Keywords: Candida albicans; drug resistance; evolution; evolutionary biology; genomics; infectious disease; microbiology; virulence.
Conflict of interest statement
AR: Senior editor,
The other authors declare that no competing interests exist.
Figures






Comment in
-
Adaptation on a genomic scale.Elife. 2015 Feb 3;4:e06193. doi: 10.7554/eLife.06193. Elife. 2015. PMID: 25647727 Free PMC article.
Similar articles
-
Genotypic evolution of azole resistance mechanisms in sequential Candida albicans isolates.Eukaryot Cell. 2007 Oct;6(10):1889-904. doi: 10.1128/EC.00151-07. Epub 2007 Aug 10. Eukaryot Cell. 2007. PMID: 17693596 Free PMC article.
-
Adaptation on a genomic scale.Elife. 2015 Feb 3;4:e06193. doi: 10.7554/eLife.06193. Elife. 2015. PMID: 25647727 Free PMC article.
-
The development of fluconazole resistance in Candida albicans - an example of microevolution of a fungal pathogen.J Microbiol. 2016 Mar;54(3):192-201. doi: 10.1007/s12275-016-5628-4. Epub 2016 Feb 27. J Microbiol. 2016. PMID: 26920879 Review.
-
Candida albicans Genetic Background Influences Mean and Heterogeneity of Drug Responses and Genome Stability during Evolution in Fluconazole.mSphere. 2020 Jun 24;5(3):e00480-20. doi: 10.1128/mSphere.00480-20. mSphere. 2020. PMID: 32581072 Free PMC article.
-
Mechanisms of genome evolution in Candida albicans.Curr Opin Microbiol. 2019 Dec;52:47-54. doi: 10.1016/j.mib.2019.05.001. Epub 2019 Jun 6. Curr Opin Microbiol. 2019. PMID: 31176092 Free PMC article. Review.
Cited by
-
Antimicrobial Peptide AMP-17 Affects Candida albicans by Disrupting Its Cell Wall and Cell Membrane Integrity.Infect Drug Resist. 2020 Jul 22;13:2509-2520. doi: 10.2147/IDR.S250278. eCollection 2020. Infect Drug Resist. 2020. PMID: 32801789 Free PMC article.
-
Evolutionary genomics of yeast pathogens in the Saccharomycotina.FEMS Yeast Res. 2016 Sep;16(6):fow064. doi: 10.1093/femsyr/fow064. Epub 2016 Aug 3. FEMS Yeast Res. 2016. PMID: 27493146 Free PMC article. Review.
-
Negative epistasis: a route to intraspecific reproductive isolation in yeast?Curr Genet. 2016 Feb;62(1):25-9. doi: 10.1007/s00294-015-0505-y. Epub 2015 Jul 12. Curr Genet. 2016. PMID: 26164016 Free PMC article. Review.
-
MycopathologiaGENOMES: The New 'Home' for the Publication of Fungal Genomes.Mycopathologia. 2019 Oct;184(5):551-554. doi: 10.1007/s11046-019-00366-3. Epub 2019 Aug 10. Mycopathologia. 2019. PMID: 31399887
-
Chemogenomic profiling to understand the antifungal action of a bioactive aurone compound.PLoS One. 2019 Dec 11;14(12):e0226068. doi: 10.1371/journal.pone.0226068. eCollection 2019. PLoS One. 2019. PMID: 31825988 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

