The R3-MYB gene GhCPC negatively regulates cotton fiber elongation
- PMID: 25646816
- PMCID: PMC4315419
- DOI: 10.1371/journal.pone.0116272
The R3-MYB gene GhCPC negatively regulates cotton fiber elongation
Abstract
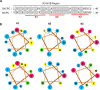
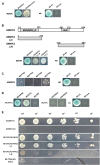
Cotton (Gossypium spp.) fibers are single-cell trichomes that arise from the outer epidermal layer of seed coat. Here, we isolated a R3-MYB gene GhCPC, identified by cDNA microarray analysis. The only conserved R3 motif and different expression between TM-1 and fuzzless-lintless mutants suggested that it might be a negative regulator in fiber development. Transgenic evidence showed that GhCPC overexpression not only delayed fiber initiation but also led to significant decreases in fiber length. Interestingly, Yeast two-hybrid analysis revealed an interaction complex, in which GhCPC and GhTTG1/4 separately interacted with GhMYC1. In transgenic plants, Q-PCR analysis showed that GhHOX3 (GL2) and GhRDL1 were significantly down regulated in -1-5 DPA ovules and fibers. In addition, Yeast one-hybrid analysis demonstrated that GhMYC1 could bind to the E-box cis-elements and the promoter of GhHOX3. These results suggested that GhHOX3 (GL2) might be downstream gene of the regulatory complex. Also, overexpression of GhCPC in tobacco led to differential loss of pigmentation. Taken together, the results suggested that GhCPC might negatively regulate cotton fiber initiation and early elongation by a potential CPC-MYC1-TTG1/4 complex. Although the fibers were shorter in transgenic cotton lines than in the wild type, no significant difference was detected in stem or leaf trichomes, even in cotton mutants (five naked seed or fuzzless), suggesting that fiber and trichome development might be regulated by two sets of genes sharing a similar model.
Conflict of interest statement
Figures






Similar articles
-
Expression profiling identifies genes expressed early during lint fibre initiation in cotton.Plant Cell Physiol. 2006 Jan;47(1):107-27. doi: 10.1093/pcp/pci228. Epub 2005 Nov 7. Plant Cell Physiol. 2006. PMID: 16278222
-
The MYB transcription factor GhMYB25 regulates early fibre and trichome development.Plant J. 2009 Jul;59(1):52-62. doi: 10.1111/j.1365-313X.2009.03847.x. Epub 2009 Feb 26. Plant J. 2009. PMID: 19309462
-
Epidermal cell differentiation in cotton mediated by the homeodomain leucine zipper gene, GhHD-1.Plant J. 2012 Aug;71(3):464-78. doi: 10.1111/j.1365-313X.2012.05003.x. Epub 2012 May 28. Plant J. 2012. PMID: 22443311
-
Gene expression changes and early events in cotton fibre development.Ann Bot. 2007 Dec;100(7):1391-401. doi: 10.1093/aob/mcm232. Epub 2007 Sep 27. Ann Bot. 2007. PMID: 17905721 Free PMC article. Review.
-
MIXTAs and phytohormones orchestrate cotton fiber development.Curr Opin Plant Biol. 2021 Feb;59:101975. doi: 10.1016/j.pbi.2020.10.007. Epub 2020 Dec 6. Curr Opin Plant Biol. 2021. PMID: 33296746 Review.
Cited by
-
Heterologous Expression of GbTCP4, a Class II TCP Transcription Factor, Regulates Trichome Formation and Root Hair Development in Arabidopsis.Genes (Basel). 2019 Sep 19;10(9):726. doi: 10.3390/genes10090726. Genes (Basel). 2019. PMID: 31546783 Free PMC article.
-
Advances in understanding epigenetic regulation of plant trichome development: a comprehensive review.Hortic Res. 2023 Jul 19;10(9):uhad145. doi: 10.1093/hr/uhad145. eCollection 2023 Sep. Hortic Res. 2023. PMID: 37691965 Free PMC article.
-
The Pivotal Role of Major Chromosomes of Sub-Genomes A and D in Fiber Quality Traits of Cotton.Front Genet. 2022 Mar 24;12:642595. doi: 10.3389/fgene.2021.642595. eCollection 2021. Front Genet. 2022. PMID: 35401652 Free PMC article. Review.
-
Plant microProteins: Small but powerful modulators of plant development.iScience. 2022 Oct 21;25(11):105400. doi: 10.1016/j.isci.2022.105400. eCollection 2022 Nov 18. iScience. 2022. PMID: 36353725 Free PMC article. Review.
-
High-density linkage map construction and QTL analyses for fiber quality, yield and morphological traits using CottonSNP63K array in upland cotton (Gossypium hirsutum L.).BMC Genomics. 2019 Nov 21;20(1):889. doi: 10.1186/s12864-019-6214-z. BMC Genomics. 2019. PMID: 31771502 Free PMC article.
References
-
- Oppenheimer DG, Herman PL, Sivakumaran S, Esch J, Marks MD (1991) A myb gene required for leaf trichome differentiation in Arabidopsis is expressed in stipules. Cell 67: 483–493. - PubMed
-
- Zhang F, Gonzalez A, Zhao M, Payne CT, Lloyd A (2003) A network of redundant bHLH proteins functions in all TTG1-dependent pathways of Arabidopsis. Development 130: 4859–4869. - PubMed
-
- Rerie WG, Feldmann KA, Marks MD (1994) The GLABRA2 gene encodes a homeo domain protein required for normal trichome development in Arabidopsis. Genes & development 8: 1388–1399. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

