Thermodynamic binding analysis of Notch transcription complexes from Drosophila melanogaster
- PMID: 25650119
- PMCID: PMC4420529
- DOI: 10.1002/pro.2652
Thermodynamic binding analysis of Notch transcription complexes from Drosophila melanogaster
Abstract
Notch is an intercellular signaling pathway that is highly conserved in metazoans and is essential for proper cellular specification during development and in the adult organism. Misregulated Notch signaling underlies or contributes to the pathogenesis of many human diseases, most notably cancer. Signaling through the Notch pathway ultimately results in changes in gene expression, which is regulated by the transcription factor CSL. Upon pathway activation, CSL forms a ternary complex with the intracellular domain of the Notch receptor (NICD) and the transcriptional coactivator Mastermind (MAM) that activates transcription from Notch target genes. While detailed in vitro studies have been conducted with mammalian and worm orthologous proteins, less is known regarding the molecular details of the Notch ternary complex in Drosophila. Here we thermodynamically characterize the assembly of the fly ternary complex using isothermal titration calorimetry. Our data reveal striking differences in the way the RAM (RBP-J associated molecule) and ANK (ankyrin) domains of NICD interact with CSL that is specific to the fly. Additional analysis using cross-species experiments suggest that these differences are primarily due to fly CSL, while experiments using point mutants show that the interface between fly CSL and ANK is likely similar to the mammalian and worm interface. Finally, we show that the binding of the fly RAM domain to CSL does not affect interactions of the corepressor Hairless with CSL. Taken together, our data suggest species-specific differences in ternary complex assembly that may be significant in understanding how CSL regulates transcription in different organisms.
Keywords: CSL; RBP-J; X-ray crystallography; isothermal titration calorimetry; notch signaling; protein−protein interactions; transcription.
© 2015 The Protein Society.
Figures

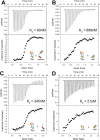
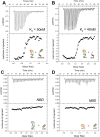
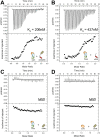
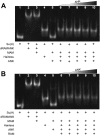
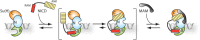
References
-
- Fortini ME. Introduction—notch in development and disease. Semin Cell Dev Biol. 2012;23:419–420. - PubMed
-
- Liu J, Sato C, Cerletti M, Wagers A. Notch signaling in the regulation of stem cell self-renewal and differentiation. Curr Top Dev Biol. 2010;92:367–409. - PubMed
-
- Radtke F, Fasnacht N, Macdonald HR. Notch signaling in the immune system. Immunity. 2010;32:14–27. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

