Genome-wide analysis of bHLH transcription factor and involvement in the infection by yellow leaf curl virus in tomato (Solanum lycopersicum)
- PMID: 25652024
- PMCID: PMC4333901
- DOI: 10.1186/s12864-015-1249-2
Genome-wide analysis of bHLH transcription factor and involvement in the infection by yellow leaf curl virus in tomato (Solanum lycopersicum)
Abstract
Background: The basic helix-loop-helix (bHLH) proteins are a superfamily of transcription factors that can bind to specific DNA target sites. They have been well characterized in model plants such as Arabidopsis and rice and have been shown to be important regulatory components in many different biological processes. However, no systemic analysis of the bHLH transcription factor family has yet been reported in tomatoes. Tomato yellow leaf curl virus (TYLCV) threatens tomato production worldwide by causing leaf yellowing, leaf curling, plant stunting and flower abscission.
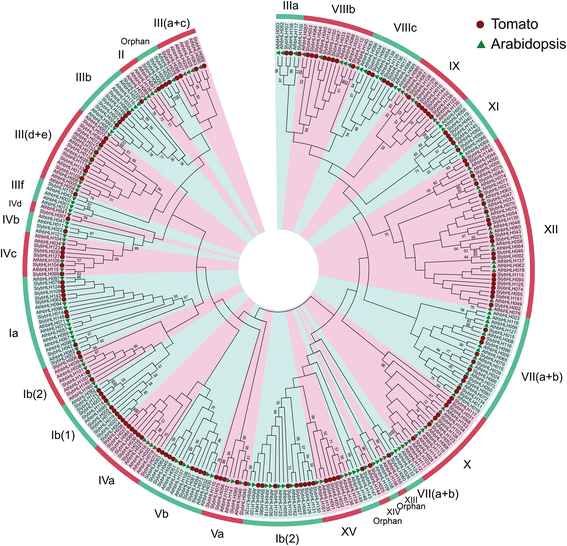
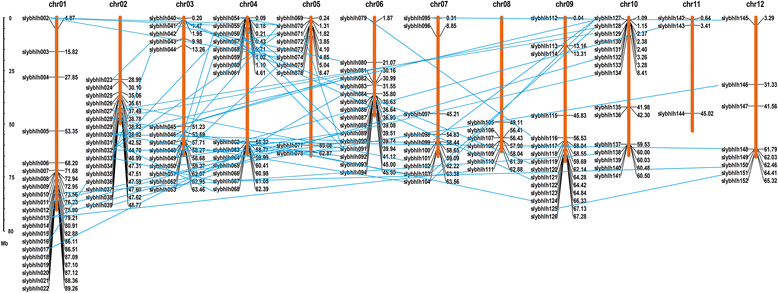
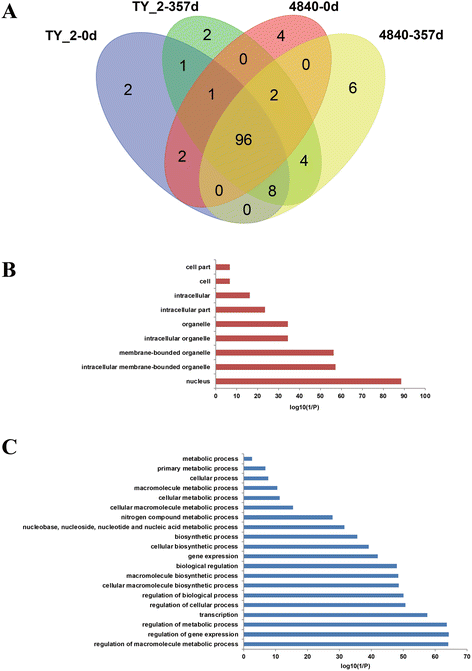
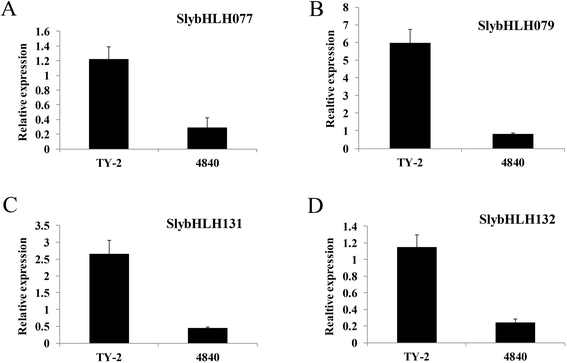
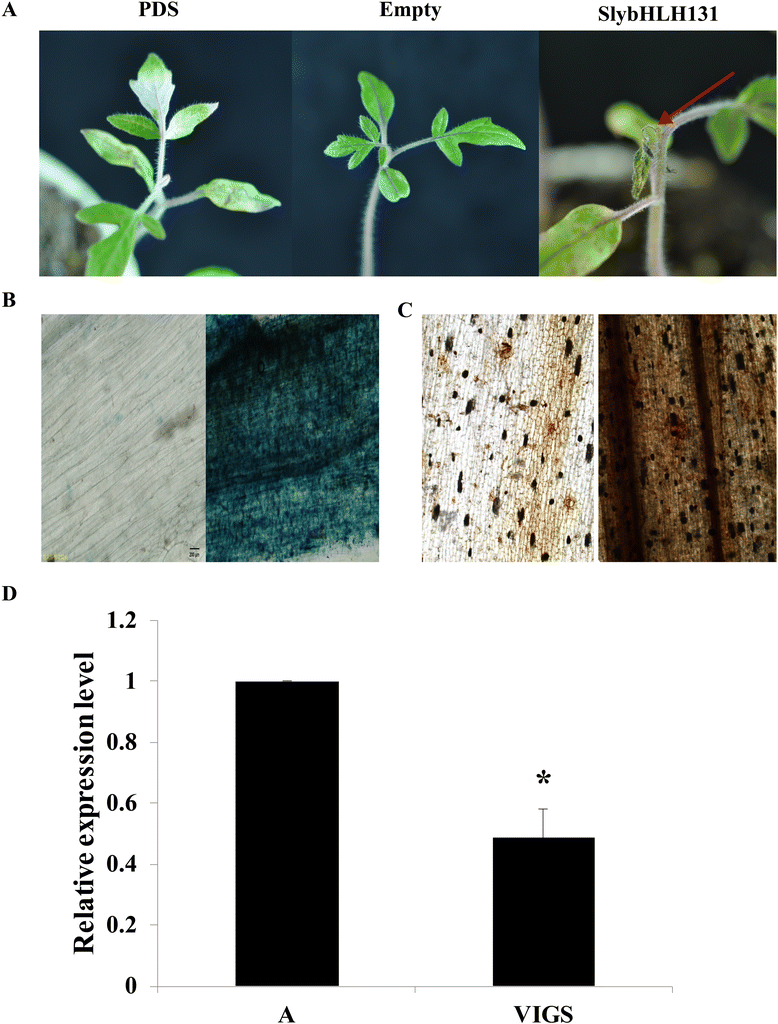
Results: A total of 152 bHLH transcription factors were identified from the entire tomato genome. Phylogenetic analysis of bHLH domain sequences from Arabidopsis and tomato facilitated classification of these genes into 26 subfamilies. The evolutionary and possible functional relationships revealed during this analysis are supported by other criteria, including the chromosomal distribution of these genes, the conservation of motifs and exon/intron structural patterns, and the predicted DNA binding activities within subfamilies. Distribution mapping results showed bHLH genes were localized on the 12 tomato chromosomes. Among the 152 bHLH genes from the tomato genome, 96 bHLH genes were detected in the TYLCV-susceptible and resistant tomato breeding line before (0 dpi) and after TYLCV (357 dpi) infection. As anticipated, gene ontology (GO) analysis indicated that most bHLH genes are related to the regulation of macromolecule metabolic processes and gene expression. Only four bHLH genes were differentially expressed between 0 and 357 dpi. Virus-induced gene silencing (VIGS) of one bHLH genes SlybHLH131 in resistant lines can lead to the cell death.
Conclusion: In the present study, 152 bHLH transcription factor genes were identified. One of which bHLH genes, SlybHLH131, was found to be involved in the TYLCV infection through qRT-PCR expression analysis and VIGS validation. The isolation and identification of these bHLH transcription factors facilitated clarification of the molecular genetic basis for the genetic improvement of tomatoes and the development of functional gene resources for transgenic research. In addition, these findings may aid in uncovering an unexplored mechanism during the TYLCV infection in tomatoes.
Figures
Similar articles
-
Comparative transcriptome profiling of a resistant vs. susceptible tomato (Solanum lycopersicum) cultivar in response to infection by tomato yellow leaf curl virus.PLoS One. 2013 Nov 18;8(11):e80816. doi: 10.1371/journal.pone.0080816. eCollection 2013. PLoS One. 2013. PMID: 24260487 Free PMC article.
-
Accumulation and transmission of alphasatellite, betasatellite and tomato yellow leaf curl virus in susceptible and Ty-1-resistant tomato plants.Virus Res. 2018 Jul 15;253:124-134. doi: 10.1016/j.virusres.2018.06.003. Epub 2018 Jun 15. Virus Res. 2018. PMID: 29908896
-
A severe symptom phenotype in tomato in Mali is caused by a reassortant between a novel recombinant begomovirus (Tomato yellow leaf curl Mali virus) and a betasatellite.Mol Plant Pathol. 2009 May;10(3):415-30. doi: 10.1111/j.1364-3703.2009.00541.x. Mol Plant Pathol. 2009. PMID: 19400843 Free PMC article.
-
Tomato yellow leaf curl virus: Characteristics, influence, and regulation mechanism.Plant Physiol Biochem. 2024 Aug;213:108812. doi: 10.1016/j.plaphy.2024.108812. Epub 2024 Jun 8. Plant Physiol Biochem. 2024. PMID: 38875781 Review.
-
Tomato Yellow Leaf Curl Virus: Impact, Challenges, and Management.Trends Plant Sci. 2020 Sep;25(9):897-911. doi: 10.1016/j.tplants.2020.03.015. Epub 2020 May 1. Trends Plant Sci. 2020. PMID: 32371058 Review.
Cited by
-
Knockout of SlMS10 Gene (Solyc02g079810) Encoding bHLH Transcription Factor Using CRISPR/Cas9 System Confers Male Sterility Phenotype in Tomato.Plants (Basel). 2020 Sep 11;9(9):1189. doi: 10.3390/plants9091189. Plants (Basel). 2020. PMID: 32933074 Free PMC article.
-
AvNAC030, a NAC Domain Transcription Factor, Enhances Salt Stress Tolerance in Kiwifruit.Int J Mol Sci. 2021 Nov 2;22(21):11897. doi: 10.3390/ijms222111897. Int J Mol Sci. 2021. PMID: 34769325 Free PMC article.
-
Genome-wide identification of the Capsicum bHLH transcription factor family: discovery of a candidate regulator involved in the regulation of species-specific bioactive metabolites.BMC Plant Biol. 2021 Jun 7;21(1):262. doi: 10.1186/s12870-021-03004-7. BMC Plant Biol. 2021. PMID: 34098881 Free PMC article.
-
Identification of Transcriptional Networks Involved in De Novo Organ Formation in Tomato Hypocotyl Explants.Int J Mol Sci. 2022 Dec 17;23(24):16112. doi: 10.3390/ijms232416112. Int J Mol Sci. 2022. PMID: 36555756 Free PMC article.
-
Analysis of bHLH genes from foxtail millet (Setaria italica) and their potential relevance to drought stress.PLoS One. 2018 Nov 9;13(11):e0207344. doi: 10.1371/journal.pone.0207344. eCollection 2018. PLoS One. 2018. PMID: 30412624 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

