GC-Content evolution in bacterial genomes: the biased gene conversion hypothesis expands
- PMID: 25659072
- PMCID: PMC4450053
- DOI: 10.1371/journal.pgen.1004941
GC-Content evolution in bacterial genomes: the biased gene conversion hypothesis expands
Abstract
The characterization of functional elements in genomes relies on the identification of the footprints of natural selection. In this quest, taking into account neutral evolutionary processes such as mutation and genetic drift is crucial because these forces can generate patterns that may obscure or mimic signatures of selection. In mammals, and probably in many eukaryotes, another such confounding factor called GC-Biased Gene Conversion (gBGC) has been documented. This mechanism generates patterns identical to what is expected under selection for higher GC-content, specifically in highly recombining genomic regions. Recent results have suggested that a mysterious selective force favouring higher GC-content exists in Bacteria but the possibility that it could be gBGC has been excluded. Here, we show that gBGC is probably at work in most if not all bacterial species. First we find a consistent positive relationship between the GC-content of a gene and evidence of intra-genic recombination throughout a broad spectrum of bacterial clades. Second, we show that the evolutionary force responsible for this pattern is acting independently from selection on codon usage, and could potentially interfere with selection in favor of optimal AU-ending codons. A comparison with data from human populations shows that the intensity of gBGC in Bacteria is comparable to what has been reported in mammals. We propose that gBGC is not restricted to sexual Eukaryotes but also widespread among Bacteria and could therefore be an ancestral feature of cellular organisms. We argue that if gBGC occurs in bacteria, it can account for previously unexplained observations, such as the apparent non-equilibrium of base substitution patterns and the heterogeneity of gene composition within bacterial genomes. Because gBGC produces patterns similar to positive selection, it is essential to take this process into account when studying the evolutionary forces at work in bacterial genomes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
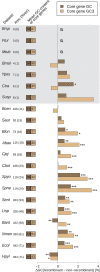


References
-
- Bernardi G, Olofsson B, Filipski J, Zerial M, Salinas J, et al. (1985) The mosaic genome of warm-blooded vertebrates. Science 228: 953–958. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

