Electrochemistry of nonconjugated proteins and glycoproteins. Toward sensors for biomedicine and glycomics
- PMID: 25659975
- PMCID: PMC4360380
- DOI: 10.1021/cr500279h
Electrochemistry of nonconjugated proteins and glycoproteins. Toward sensors for biomedicine and glycomics
Figures
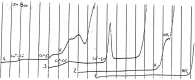
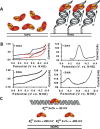
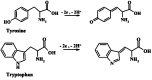


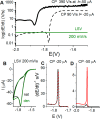
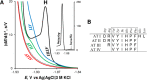

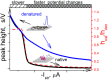
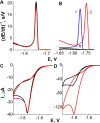
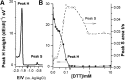


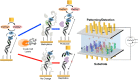
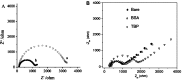
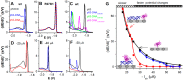

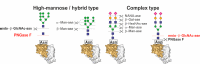

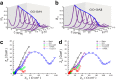
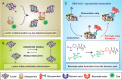
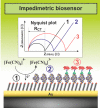

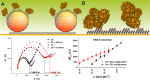
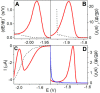

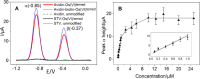
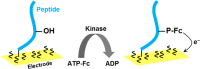
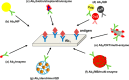

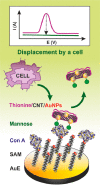
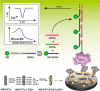
Similar articles
-
Recent Progress in Electrochemical Biosensors for Glycoproteins.Sensors (Basel). 2016 Dec 1;16(12):2045. doi: 10.3390/s16122045. Sensors (Basel). 2016. PMID: 27916961 Free PMC article. Review.
-
Electrochemical sensors and biosensors.Anal Chem. 2012 Jan 17;84(2):685-707. doi: 10.1021/ac202878q. Epub 2011 Nov 11. Anal Chem. 2012. PMID: 22044045 Free PMC article. Review. No abstract available.
-
Electrochemical biosensors on platforms of graphene.Chem Commun (Camb). 2013 Oct 25;49(83):9526-39. doi: 10.1039/c3cc44735a. Chem Commun (Camb). 2013. PMID: 24025792 Review.
-
Electrochemical sensors and biosensors based on nanomaterials and nanostructures.Anal Chem. 2015 Jan 6;87(1):230-49. doi: 10.1021/ac5039863. Epub 2014 Dec 19. Anal Chem. 2015. PMID: 25354297 Free PMC article. Review. No abstract available.
-
New electrochemical methods.Anal Chem. 2012 Jan 17;84(2):669-84. doi: 10.1021/ac2026767. Epub 2011 Nov 9. Anal Chem. 2012. PMID: 22017638 Review. No abstract available.
Cited by
-
Full-length antibodies versus single-chain antibody fragments for a selective impedimetric lectin-based glycoprofiling of prostate specific antigen.Electrochim Acta. 2017 Aug 20;246:399-405. doi: 10.1016/j.electacta.2017.06.065. Electrochim Acta. 2017. PMID: 29104305 Free PMC article.
-
Biosensors in Health Care: The Milestones Achieved in Their Development towards Lab-on-Chip-Analysis.Biochem Res Int. 2016;2016:3130469. doi: 10.1155/2016/3130469. Epub 2016 Mar 3. Biochem Res Int. 2016. PMID: 27042353 Free PMC article. Review.
-
Electrochemical detection of different p53 conformations by using nanostructured surfaces.Sci Rep. 2019 Nov 22;9(1):17347. doi: 10.1038/s41598-019-53994-6. Sci Rep. 2019. PMID: 31758050 Free PMC article.
-
Aberrant sialylation of a prostate-specific antigen: Electrochemical label-free glycoprofiling in prostate cancer serum samples.Anal Chim Acta. 2016 Aug 31;934:72-9. doi: 10.1016/j.aca.2016.06.043. Epub 2016 Jul 1. Anal Chim Acta. 2016. PMID: 27506346 Free PMC article.
-
Graphene as a signal amplifier for preparation of ultrasensitive electrochemical biosensors.Chem Zvesti. 2015 Jan;69(1):112-133. doi: 10.1515/chempap-2015-0051. Epub 2014 Nov 28. Chem Zvesti. 2015. PMID: 27242391 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

