Crystal structure of the Rad3/XPD regulatory domain of Ssl1/p44
- PMID: 25681444
- PMCID: PMC4375486
- DOI: 10.1074/jbc.M115.636514
Crystal structure of the Rad3/XPD regulatory domain of Ssl1/p44
Abstract
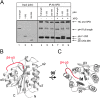
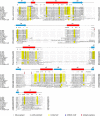
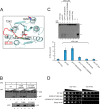
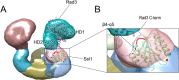
The Ssl1/p44 subunit is a core component of the yeast/mammalian general transcription factor TFIIH, which is involved in transcription and DNA repair. Ssl1/p44 binds to and stimulates the Rad3/XPD helicase activity of TFIIH. To understand the helicase stimulatory mechanism of Ssl1/p44, we determined the crystal structure of the N-terminal regulatory domain of Ssl1 from Saccharomyces cerevisiae. Ssl1 forms a von Willebrand factor A fold in which a central six-stranded β-sheet is sandwiched between three α helices on both sides. Structural and biochemical analyses of Ssl1/p44 revealed that the β4-α5 loop, which is frequently found at the interface between von Willebrand factor A family proteins and cellular counterparts, is critical for the stimulation of Rad3/XPD. Yeast genetics analyses showed that double mutation of Leu-239 and Ser-240 in the β4-α5 loop of Ssl1 leads to lethality of a yeast strain, demonstrating the importance of the Rad3-Ssl1 interactions to cell viability. Here, we provide a structural model for the Rad3/XPD-Ssl1/p44 complex and insights into how the binding of Ssl1/p44 contributes to the helicase activity of Rad3/XPD and cell viability.
Keywords: Crystal Structure; DNA Repair; Nucleotide Excision Repair; Rad3/XPD Helicase; Regulatory Domain of Ssl1/p44; TFIIH; Transcription; Yeast Genetics.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


Similar articles
-
p44/SSL1, the regulatory subunit of the XPD/RAD3 helicase, plays a crucial role in the transcriptional activity of TFIIH.J Biol Chem. 2000 Oct 27;275(43):33260-6. doi: 10.1074/jbc.M004764200. J Biol Chem. 2000. PMID: 10924514
-
Arabidopsis homologue of human transcription factor IIH/nucleotide excision repair factor p44 can function in transcription and DNA repair and interacts with AtXPD.Plant J. 2006 May;46(3):512-21. doi: 10.1111/j.1365-313X.2006.02705.x. Plant J. 2006. PMID: 16623910
-
Architecture of the Human and Yeast General Transcription and DNA Repair Factor TFIIH.Mol Cell. 2015 Sep 3;59(5):794-806. doi: 10.1016/j.molcel.2015.07.016. Mol Cell. 2015. PMID: 26340423 Free PMC article.
-
The long unwinding road: XPB and XPD helicases in damaged DNA opening.Cell Cycle. 2010 Jan 1;9(1):90-6. doi: 10.4161/cc.9.1.10267. Epub 2010 Jan 5. Cell Cycle. 2010. PMID: 20016270 Review.
-
DNA helicases, genomic instability, and human genetic disease.Annu Rev Genomics Hum Genet. 2000;1:409-59. doi: 10.1146/annurev.genom.1.1.409. Annu Rev Genomics Hum Genet. 2000. PMID: 11701636 Review.
Cited by
-
The complete structure of the human TFIIH core complex.Elife. 2019 Mar 12;8:e44771. doi: 10.7554/eLife.44771. Elife. 2019. PMID: 30860024 Free PMC article.
-
RecQ and Fe-S helicases have unique roles in DNA metabolism dictated by their unwinding directionality, substrate specificity, and protein interactions.Biochem Soc Trans. 2018 Feb 19;46(1):77-95. doi: 10.1042/BST20170044. Epub 2017 Dec 22. Biochem Soc Trans. 2018. PMID: 29273621 Free PMC article. Review.
-
Near-atomic resolution visualization of human transcription promoter opening.Nature. 2016 May 19;533(7603):359-65. doi: 10.1038/nature17970. Epub 2016 May 11. Nature. 2016. PMID: 27193682 Free PMC article.
-
The Structures of Eukaryotic Transcription Pre-initiation Complexes and Their Functional Implications.Subcell Biochem. 2019;93:143-192. doi: 10.1007/978-3-030-28151-9_5. Subcell Biochem. 2019. PMID: 31939151 Free PMC article. Review.
-
The intricate network between the p34 and p44 subunits is central to the activity of the transcription/DNA repair factor TFIIH.Nucleic Acids Res. 2017 Oct 13;45(18):10872-10883. doi: 10.1093/nar/gkx743. Nucleic Acids Res. 2017. PMID: 28977422 Free PMC article.
References
-
- Kim T. K., Ebright R. H., Reinberg D. (2000) Mechanism of ATP-dependent promoter melting by transcription factor IIH. Science 288, 1418–1422 - PubMed
-
- Compe E., Egly J. M. (2012) TFIIH: when transcription met DNA repair. Nat. Rev. Mol. Cell Biol. 13, 343–354 - PubMed
-
- Schultz P., Fribourg S., Poterszman A., Mallouh V., Moras D., Egly J. M. (2000) Molecular structure of human TFIIH. Cell 102, 599–607 - PubMed
-
- Serizawa H., Conaway J. W., Conaway R. C. (1993) Phosphorylation of C-terminal domain of RNA polymerase II is not required in basal transcription. Nature 363, 371–374 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

