Non-allelic gene conversion enables rapid evolutionary change at multiple regulatory sites encoded by transposable elements
- PMID: 25688566
- PMCID: PMC4384637
- DOI: 10.7554/eLife.05899
Non-allelic gene conversion enables rapid evolutionary change at multiple regulatory sites encoded by transposable elements
Abstract
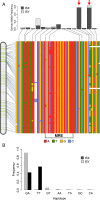
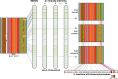
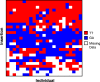
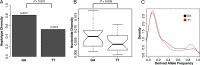
Transposable elements (TEs) allow rewiring of regulatory networks, and the recent amplification of the ISX element dispersed 77 functional but suboptimal binding sites for the dosage compensation complex to a newly formed X chromosome in Drosophila. Here we identify two linked refining mutations within ISX that interact epistatically to increase binding affinity to the dosage compensation complex. Selection has increased the frequency of this derived haplotype in the population, which is fixed at 30% of ISX insertions and polymorphic among another 41%. Sharing of this haplotype indicates that high levels of gene conversion among ISX elements allow them to 'crowd-source' refining mutations, and a refining mutation that occurs at any single ISX element can spread in two dimensions: horizontally across insertion sites by non-allelic gene conversion, and vertically through the population by natural selection. These results describe a novel route by which fully functional regulatory elements can arise rapidly from TEs and implicate non-allelic gene conversion as having an important role in accelerating the evolutionary fine-tuning of regulatory networks.
Keywords: Drosophila miranda; evolutionary biology; genomics; non-allelic gene conversion; regulatory networks; transposable elements.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures





Comment in
-
Spreading good news.Elife. 2015 Apr 2;4:e07108. doi: 10.7554/eLife.07108. Elife. 2015. PMID: 25838131 Free PMC article.
References
-
- Alekseyenko AA, Ellison CE, Gorchakov AA, Zhou Q, Kaiser VB, Toda N, Walton Z, Peng S, Park PJ, Bachtrog D, Kuroda MI. Conservation and de novo acquisition of dosage compensation on newly evolved sex chromosomes in Drosophila. Genes & Development. 2013;27:853–858. doi: 10.1101/gad.215426.113. - DOI - PMC - PubMed
-
- Alekseyenko AA, Peng S, Larschan E, Gorchakov AA, Lee OK, Kharchenko P, McGrath SD, Wang CI, Mardis ER, Park PJ, Kuroda MI. A sequence motif within chromatin entry sites directs MSL establishment on the Drosophila X chromosome. Cell. 2008;134:599–609. doi: 10.1016/j.cell.2008.06.033. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

