Role of epigenetic modifications in luminal breast cancer
- PMID: 25689414
- PMCID: PMC4539290
- DOI: 10.2217/epi.15.10
Role of epigenetic modifications in luminal breast cancer
Abstract
Luminal breast cancers represent approximately 75% of cases. Explanations into the causes of endocrine resistance are complex and are generally ascribed to genomic mechanisms. Recently, attention has been drawn to the role of epigenetic modifications in hormone resistance. We review these here. Epigenetic modifications are reversible, heritable and include changes in DNA methylation patterns, modification of histones and altered microRNA expression levels that target the receptors or their signaling pathways. Large-scale analyses indicate distinct epigenomic profiles that distinguish breast cancers from normal and benign tissues. Taking advantage of the reversibility of epigenetic modifications, drugs that target epigenetic modifiers, given in combination with chemotherapies or endocrine therapies, may represent promising approaches to restoration of therapy responsiveness in these cases.
Keywords: breast cancer; epigenetic modifications; estrogen receptors; progesterone receptors.
Conflict of interest statement
The authors are grateful for support from the NIH (RO1-CA026869-35), the Breast Cancer Research Foundation and the National Foundation for Cancer Research. The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
No writing assistance was utilized in the production of this manuscript.
Figures
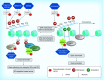
References
-
- Prat A, Karginova O, Parker JS, et al. Characterization of cell lines derived from breast cancers and normal mammary tissues for the study of the intrinsic molecular subtypes. Breast Cancer Res. Treat. 2013;142(2):237–255. - PMC - PubMed
-
•• Data presented in this paper help to improve our understanding of the wide range of breast cancer cell line models through the appropriate pairing of cell lines with relevant in vivo tumor and normal cell counterpart.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
