Characterization of a large cluster of influenza A(H1N1)pdm09 viruses cross-resistant to oseltamivir and peramivir during the 2013-2014 influenza season in Japan
- PMID: 25691635
- PMCID: PMC4394804
- DOI: 10.1128/AAC.04836-14
Characterization of a large cluster of influenza A(H1N1)pdm09 viruses cross-resistant to oseltamivir and peramivir during the 2013-2014 influenza season in Japan
Abstract
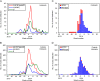
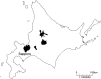
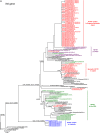
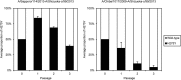
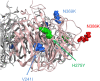
Between September 2013 and July 2014, 2,482 influenza 2009 pandemic A(H1N1) [A(H1N1)pdm09] viruses were screened in Japan for the H275Y substitution in their neuraminidase (NA) protein, which confers cross-resistance to oseltamivir and peramivir. We found that a large cluster of the H275Y mutant virus was present prior to the main influenza season in Sapporo /: Hokkaido, with the detection rate for this mutant virus reaching 29% in this area. Phylogenetic analysis suggested the clonal expansion of a single mutant virus in Sapporo /: Hokkaido. To understand the reason for this large cluster, we examined the in vitro and in vivo properties of the mutant virus. We found that it grew well in cell culture, with growth comparable to that of the wild-type virus. The cluster virus also replicated well in the upper respiratory tract of ferrets and was transmitted efficiently between ferrets by way of respiratory droplets. Almost all recently circulating A(H1N1)pdm09 viruses, including the cluster virus, possessed two substitutions in NA, V241I and N369K, which are known to increase replication and transmission fitness. A structural analysis of NA predicted that a third substitution (N386K) in the NA of the cluster virus destabilized the mutant NA structure in the presence of the V241I and N369K substitutions. Our results suggest that the cluster virus retained viral fitness to spread among humans and, accordingly, caused the large cluster in Sapporo/Hokkaido. However, the mutant NA structure was less stable than that of the wild-type virus. Therefore, once the wild-type virus began to circulate in the community, the mutant virus could not compete and faded out.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures


Similar articles
-
Characteristics of oseltamivir-resistant influenza A (H1N1) pdm09 virus during the 2013-2014 influenza season in Mainland China.Virol J. 2015 Jun 24;12:96. doi: 10.1186/s12985-015-0317-1. Virol J. 2015. PMID: 26103966 Free PMC article.
-
Characterization of neuraminidase inhibitor-resistant influenza A(H1N1)pdm09 viruses isolated in four seasons during pandemic and post-pandemic periods in Japan.Influenza Other Respir Viruses. 2013 Nov;7(6):1390-9. doi: 10.1111/irv.12132. Epub 2013 Jun 8. Influenza Other Respir Viruses. 2013. PMID: 23745712 Free PMC article.
-
Impact of potential permissive neuraminidase mutations on viral fitness of the H275Y oseltamivir-resistant influenza A(H1N1)pdm09 virus in vitro, in mice and in ferrets.J Virol. 2014 Feb;88(3):1652-8. doi: 10.1128/JVI.02681-13. Epub 2013 Nov 20. J Virol. 2014. PMID: 24257597 Free PMC article.
-
[Current anti-influenza virus chemotherapy].Nihon Rinsho. 2010 Sep;68(9):1679-84. Nihon Rinsho. 2010. PMID: 20845747 Review. Japanese.
-
The epidemiology and spread of drug resistant human influenza viruses.Curr Opin Virol. 2014 Oct;8:22-9. doi: 10.1016/j.coviro.2014.04.009. Epub 2014 May 24. Curr Opin Virol. 2014. PMID: 24866471 Review.
Cited by
-
Antiviral options and therapeutics against influenza: history, latest developments and future prospects.Front Cell Infect Microbiol. 2023 Nov 29;13:1269344. doi: 10.3389/fcimb.2023.1269344. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 38094741 Free PMC article. Review.
-
Influenza Virus Neuraminidase Structure and Functions.Front Microbiol. 2019 Jan 29;10:39. doi: 10.3389/fmicb.2019.00039. eCollection 2019. Front Microbiol. 2019. PMID: 30761095 Free PMC article. Review.
-
Influenza A(H1N1)pdm09 Virus with Reduced Susceptibility to Baloxavir, Japan, 2024.Emerg Infect Dis. 2025 May;31(5):1019-1023. doi: 10.3201/eid3105.241123. Epub 2025 Apr 3. Emerg Infect Dis. 2025. PMID: 40180581 Free PMC article.
-
C646, a Novel p300/CREB-Binding Protein-Specific Inhibitor of Histone Acetyltransferase, Attenuates Influenza A Virus Infection.Antimicrob Agents Chemother. 2015 Dec 28;60(3):1902-6. doi: 10.1128/AAC.02055-15. Antimicrob Agents Chemother. 2015. PMID: 26711748 Free PMC article.
-
Monitoring the fitness of antiviral-resistant influenza strains during an epidemic: a mathematical modelling study.Lancet Infect Dis. 2017 Mar;17(3):339-347. doi: 10.1016/S1473-3099(16)30465-0. Epub 2016 Dec 1. Lancet Infect Dis. 2017. PMID: 27914853 Free PMC article.
References
-
- World Health Organization. 2010. WHO guidelines for pharmacological management of pandemic influenza A(H1N1) 2009 and other influenza viruses. World Health Organization, Geneva, Switzerland: http://www.who.int/csr/resources/publications/swineflu/h1n1_use_antivira.... - PubMed
-
- Monto AS, McKimm-Breschkin JL, Macken C, Hampson AW, Hay A, Klimov A, Tashiro M, Webster RG, Aymard M, Hayden FG, Zambon M. 2006. Detection of influenza viruses resistant to neuraminidase inhibitors in global surveillance during the first 3 years of their use. Antimicrob Agents Chemother 50:2395–2402. doi:10.1128/AAC.01339-05. - DOI - PMC - PubMed
-
- Ujike M, Shimabukuro K, Mochizuki K, Obuchi M, Kageyama T, Shirakura M, Kishida N, Yamashita K, Horikawa H, Kato Y, Fujita N, Tashiro M, Odagiri T, Working Group for Influenza Virus Surveillance in Japan. 2010. Oseltamivir-resistant influenza viruses A(H1N1) during 2007-2009 influenza seasons, Japan. Emerg Infect Dis 16:926–935. doi:10.3201/eid1606.091623. - DOI - PMC - PubMed
-
- Ujike M, Ejima M, Anraku A, Shimabukuro K, Obuchi M, Kishida N, Hong X, Takashita E, Fujisaki S, Yamashita K, Horikawa H, Kato Y, Oguchi A, Fujita N, Tashiro M, Odagiri T, Influenza Virus Surveillance Group of Japan. 2011. Monitoring and characterization of oseltamivir-resistant pandemic (H1N1) 2009 virus, Japan, 2009–2010. Emerg Infect Dis 17:470–479. doi:10.3201/eid1703.101188. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

