Whole cell formaldehyde cross-linking simplifies purification of mitochondrial nucleoids and associated proteins involved in mitochondrial gene expression
- PMID: 25695250
- PMCID: PMC4335056
- DOI: 10.1371/journal.pone.0116726
Whole cell formaldehyde cross-linking simplifies purification of mitochondrial nucleoids and associated proteins involved in mitochondrial gene expression
Abstract
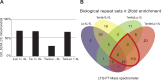
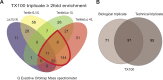
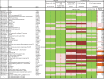
Mitochondrial DNA/protein complexes (nucleoids) appear as discrete entities inside the mitochondrial network when observed by live-cell imaging and immunofluorescence. This somewhat trivial observation in recent years has spurred research towards isolation of these complexes and the identification of nucleoid-associated proteins. Here we show that whole cell formaldehyde crosslinking combined with affinity purification and tandem mass-spectrometry provides a simple and reproducible method to identify potential nucleoid associated proteins. The method avoids spurious mitochondrial isolation and subsequent multifarious nucleoid enrichment protocols and can be implemented to allow for label-free quantification (LFQ) by mass-spectrometry. Using expression of a Flag-tagged Twinkle helicase and appropriate controls we show that this method identifies many previously identified nucleoid associated proteins. Using LFQ to compare HEK293 cells with and without mtDNA, but both expressing Twinkle-FLAG, identifies many proteins that are reduced or absent in the absence of mtDNA. This set not only includes established mtDNA maintenance proteins but also many proteins involved in mitochondrial RNA metabolism and translation and therefore represents what can be considered an mtDNA gene expression proteome. Our data provides a very valuable resource for both basic mitochondrial researchers as well as clinical geneticists working to identify novel disease genes on the basis of exome sequence data.
Conflict of interest statement
Figures


References
-
- Schatz G, Tuppy H, Haslbrun E (1964) Deoxyribonucleic Acid Associated with Yeast Mitochondria. Biochem Biophys Res Commun 15: 127–132. - PubMed
-
- Nass MM (1969) Mitochondrial DNA. I. Intramitochondrial distribution and structural relations of single- and double-length circular DNA. J Mol Biol 42: 521–528. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

