Co-evolution of Mycobacterium tuberculosis and Homo sapiens
- PMID: 25703549
- PMCID: PMC4339235
- DOI: 10.1111/imr.12264
Co-evolution of Mycobacterium tuberculosis and Homo sapiens
Abstract
The causative agent of human tuberculosis (TB), Mycobacterium tuberculosis, is an obligate pathogen that evolved to exclusively persist in human populations. For M. tuberculosis to transmit from person to person, it has to cause pulmonary disease. Therefore, M. tuberculosis virulence has likely been a significant determinant of the association between M. tuberculosis and humans. Indeed, the evolutionary success of some M. tuberculosis genotypes seems at least partially attributable to their increased virulence. The latter possibly evolved as a consequence of human demographic expansions. If co-evolution occurred, humans would have counteracted to minimize the deleterious effects of M. tuberculosis virulence. The fact that human resistance to infection has a strong genetic basis is a likely consequence of such a counter-response. The genetic architecture underlying human resistance to M. tuberculosis remains largely elusive. However, interactions between human genetic polymorphisms and M. tuberculosis genotypes have been reported. Such interactions are consistent with local adaptation and allow for a better understanding of protective immunity in TB. Future 'genome-to-genome' studies, in which locally associated human and M. tuberculosis genotypes are interrogated in conjunction, will help identify new protective antigens for the development of better TB vaccines.
Keywords: adaptation; host; pathogen; selection; virulence.
© 2015 The Authors. Immunological Reviews published by John Wiley & Sons Ltd.
Figures
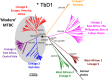




References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

