A proposal: Evolution of PCNA's role as a marker of newly replicated DNA
- PMID: 25704660
- PMCID: PMC4426064
- DOI: 10.1016/j.dnarep.2015.01.015
A proposal: Evolution of PCNA's role as a marker of newly replicated DNA
Abstract
Processivity clamps that hold DNA polymerases to DNA for processivity were the first proteins known to encircle the DNA duplex. At the time, polymerase processivity was thought to be the only function of ring shaped processivity clamps. But studies from many laboratories have identified numerous proteins that bind and function with sliding clamps. Among these processes are mismatch repair and nucleosome assembly. Interestingly, there exist polymerases that are highly processive and do not require clamps. Hence, DNA polymerase processivity does not intrinsically require that sliding clamps evolved for this purpose. We propose that polymerases evolved to require clamps as a way of ensuring that clamps are deposited on newly replicated DNA. These clamps are then used on the newly replicated daughter strands, for processes important to genomic integrity, such as mismatch repair and the assembly of nucleosomes to maintain epigenetic states of replicating cells during development.
Keywords: DNA replication; PCNA; β-Clamp.
Copyright © 2015 Elsevier B.V. All rights reserved.
Figures
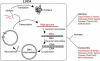




Similar articles
-
Clamp loading, unloading and intrinsic stability of the PCNA, beta and gp45 sliding clamps of human, E. coli and T4 replicases.Genes Cells. 1996 Jan;1(1):101-13. doi: 10.1046/j.1365-2443.1996.07007.x. Genes Cells. 1996. PMID: 9078370
-
Water skating: How polymerase sliding clamps move on DNA.FEBS J. 2021 Dec;288(24):7256-7262. doi: 10.1111/febs.15740. Epub 2021 Feb 18. FEBS J. 2021. PMID: 33523561 Free PMC article. Review.
-
The ring-type polymerase sliding clamp family.Genome Biol. 2001;2(1):REVIEWS3001. doi: 10.1186/gb-2001-2-1-reviews3001. Epub 2001 Jan 9. Genome Biol. 2001. PMID: 11178284 Free PMC article. Review.
-
Cellular DNA replicases: components and dynamics at the replication fork.Annu Rev Biochem. 2005;74:283-315. doi: 10.1146/annurev.biochem.73.011303.073859. Annu Rev Biochem. 2005. PMID: 15952889 Review.
-
Structural and functional similarities of prokaryotic and eukaryotic DNA polymerase sliding clamps.Nucleic Acids Res. 1995 Sep 25;23(18):3613-20. doi: 10.1093/nar/23.18.3613. Nucleic Acids Res. 1995. PMID: 7478986 Free PMC article. Review.
Cited by
-
A Replisome's journey through the bacterial chromosome.Front Microbiol. 2015 Jun 5;6:562. doi: 10.3389/fmicb.2015.00562. eCollection 2015. Front Microbiol. 2015. PMID: 26097470 Free PMC article. Review.
-
Evolution of the methyl directed mismatch repair system in Escherichia coli.DNA Repair (Amst). 2016 Feb;38:32-41. doi: 10.1016/j.dnarep.2015.11.016. Epub 2015 Dec 2. DNA Repair (Amst). 2016. PMID: 26698649 Free PMC article. Review.
-
From Processivity to Genome Maintenance: The Many Roles of Sliding Clamps.Genes (Basel). 2022 Nov 7;13(11):2058. doi: 10.3390/genes13112058. Genes (Basel). 2022. PMID: 36360296 Free PMC article. Review.
-
DNA Replication: How Does a Sliding Clamp Slide?Curr Biol. 2017 Mar 6;27(5):R174-R176. doi: 10.1016/j.cub.2017.01.053. Curr Biol. 2017. PMID: 28267969 Free PMC article.
-
Prime Editing in Dividing and Quiescent Cells.Int J Mol Sci. 2025 Apr 11;26(8):3596. doi: 10.3390/ijms26083596. Int J Mol Sci. 2025. PMID: 40332080 Free PMC article. Review.
References
-
- Watson JD, Crick FH. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature. 1953;171:737–738. http://dx.doi.org/10.1038/171738a0. - DOI - PubMed
-
- Watson JD, Crick FH. Genetical implications of the structure of deoxyribonucleic acid. Nature. 1953;171:964–967. http://dx.doi.org/10.1038/171964b0. - DOI - PubMed
-
- Lehman IR, Bessman MJ, Simms ES, Kornberg A. Enzymatic synthesis of deoxyribonucleic acid. I. Preparation of substrates and partial purification of an enzyme from Escherichia coli. J Biol Chem. 1958;233:163–170. - PubMed
-
- Kornberg A, Baker TA. DNA Replication. 2nd ed. University Science Books; New York: 2005.
-
- Kelch BA, Makino DL, O'Donnell M, Kuriyan J. Clamp loader ATPases and the evolution of DNA replication machinery. BMC Biol. 2012;10:34. http://dx.doi.org/10.1186/1741-7007-10-34. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

