A multiancestral genome-wide exome array study of Alzheimer disease, frontotemporal dementia, and progressive supranuclear palsy
- PMID: 25706306
- PMCID: PMC4397175
- DOI: 10.1001/jamaneurol.2014.4040
A multiancestral genome-wide exome array study of Alzheimer disease, frontotemporal dementia, and progressive supranuclear palsy
Abstract
Importance: Previous studies have indicated a heritable component of the etiology of neurodegenerative diseases such as Alzheimer disease (AD), frontotemporal dementia (FTD), and progressive supranuclear palsy (PSP). However, few have examined the contribution of low-frequency coding variants on a genome-wide level.
Objective: To identify low-frequency coding variants that affect susceptibility to AD, FTD, and PSP.
Design, setting, and participants: We used the Illumina HumanExome BeadChip array to genotype a large number of variants (most of which are low-frequency coding variants) in a cohort of patients with neurodegenerative disease (224 with AD, 168 with FTD, and 48 with PSP) and in 224 control individuals without dementia enrolled between 2005-2012 from multiple centers participating in the Genetic Investigation in Frontotemporal Dementia and Alzheimer's Disease (GIFT) Study. An additional multiancestral replication cohort of 240 patients with AD and 240 controls without dementia was used to validate suggestive findings. Variant-level association testing and gene-based testing were performed.
Main outcomes and measures: Statistical association of genetic variants with clinical diagnosis of AD, FTD, and PSP.
Results: Genetic variants typed by the exome array explained 44%, 53%, and 57% of the total phenotypic variance of AD, FTD, and PSP, respectively. An association with the known AD gene ABCA7 was replicated in several ancestries (discovery P=.0049, European P=.041, African American P=.043, and Asian P=.027), suggesting that exonic variants within this gene modify AD susceptibility. In addition, 2 suggestive candidate genes, DYSF (P=5.53×10(-5)) and PAXIP1 (P=2.26×10(-4)), were highlighted in patients with AD and differentially expressed in AD brain. Corroborating evidence from other exome array studies and gene expression data points toward potential involvement of these genes in the pathogenesis of AD.
Conclusions and relevance: Low-frequency coding variants with intermediate effect size may account for a significant fraction of the genetic susceptibility to AD and FTD. Furthermore, we found evidence that coding variants in the known susceptibility gene ABCA7, as well as candidate genes DYSF and PAXIP1, confer risk for AD.
Conflict of interest statement
Figures

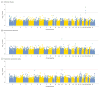
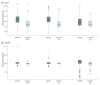
Comment in
-
Probing the exome in Alzheimer disease and other neurodegenerative disorders.JAMA Neurol. 2015 Apr;72(4):389-91. doi: 10.1001/jamaneurol.2014.4495. JAMA Neurol. 2015. PMID: 25706056 No abstract available.
References
-
- Lendon CL, Ashall F, Goate AM. Exploring the etiology of Alzheimer disease using molecular genetics. JAMA. 1997;277(10):825–831. - PubMed
-
- Lambert JC, Ibrahim-Verbaas CA, Harold D, et al. European Alzheimer’s Disease Initiative (EADI); Genetic and Environmental Risk in Alzheimer’s Disease; Alzheimer’s Disease Genetic Consortium; Cohorts for Heart and Aging Research in Genomic Epidemiology. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45(12):1452–1458. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
- RC1 AG035610/AG/NIA NIH HHS/United States
- P50 AG023501/AG/NIA NIH HHS/United States
- R01 AG041762/AG/NIA NIH HHS/United States
- P50 AG16570/AG/NIA NIH HHS/United States
- P50 AG005142/AG/NIA NIH HHS/United States
- U24 AG021886/AG/NIA NIH HHS/United States
- P50 AG05142/AG/NIA NIH HHS/United States
- R01 AG042292/AG/NIA NIH HHS/United States
- R01 AG038791/AG/NIA NIH HHS/United States
- P01 AG000538/AG/NIA NIH HHS/United States
- U01 AG016979/AG/NIA NIH HHS/United States
- U01 AG016976/AG/NIA NIH HHS/United States
- P30 NS062691/NS/NINDS NIH HHS/United States
- UL1 TR001108/TR/NCATS NIH HHS/United States
- P50 AG016573/AG/NIA NIH HHS/United States
- P01 AG019724/AG/NIA NIH HHS/United States
- P50 AG016570/AG/NIA NIH HHS/United States
- P0I AG000538/AG/NIA NIH HHS/United States
- R01 AG032306/AG/NIA NIH HHS/United States
- P50 AG025688/AG/NIA NIH HHS/United States
- R01 AG26938/AG/NIA NIH HHS/United States
- F31 NS084556/NS/NINDS NIH HHS/United States
- UL1 TR001414/TR/NCATS NIH HHS/United States
- R01 MH097268/MH/NIMH NIH HHS/United States
- P30 AG010129/AG/NIA NIH HHS/United States
- U54 NS092089/NS/NINDS NIH HHS/United States
- U24 AG21886/AG/NIA NIH HHS/United States
- R01 AG026938/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

