Protein sectors: statistical coupling analysis versus conservation
- PMID: 25723535
- PMCID: PMC4344308
- DOI: 10.1371/journal.pcbi.1004091
Protein sectors: statistical coupling analysis versus conservation
Abstract
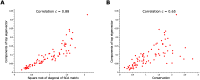
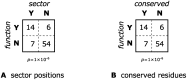
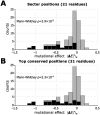
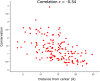
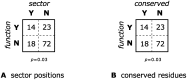
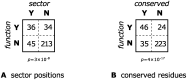
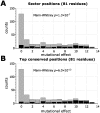
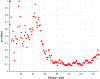
Statistical coupling analysis (SCA) is a method for analyzing multiple sequence alignments that was used to identify groups of coevolving residues termed "sectors". The method applies spectral analysis to a matrix obtained by combining correlation information with sequence conservation. It has been asserted that the protein sectors identified by SCA are functionally significant, with different sectors controlling different biochemical properties of the protein. Here we reconsider the available experimental data and note that it involves almost exclusively proteins with a single sector. We show that in this case sequence conservation is the dominating factor in SCA, and can alone be used to make statistically equivalent functional predictions. Therefore, we suggest shifting the experimental focus to proteins for which SCA identifies several sectors. Correlations in protein alignments, which have been shown to be informative in a number of independent studies, would then be less dominated by sequence conservation.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Do CB, Katoh K (2008) Protein multiple sequence alignment. Methods in Molecular Biology 484: 379–413. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

