From Protease to Decarboxylase: THE MOLECULAR METAMORPHOSIS OF PHOSPHATIDYLSERINE DECARBOXYLASE
- PMID: 25724650
- PMCID: PMC4409258
- DOI: 10.1074/jbc.M115.642413
From Protease to Decarboxylase: THE MOLECULAR METAMORPHOSIS OF PHOSPHATIDYLSERINE DECARBOXYLASE
Abstract
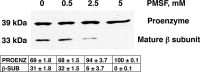
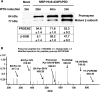
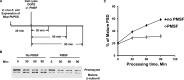
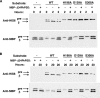
Phosphatidylserine decarboxylase (PSDs) play a central role in the synthesis of phosphatidylethanolamine in numerous species of prokaryotes and eukaryotes. PSDs are unusual decarboxylase containing a pyruvoyl prosthetic group within the active site. The covalently attached pyruvoyl moiety is formed in a concerted reaction when the PSD proenzyme undergoes an endoproteolytic cleavage into a large β-subunit, and a smaller α-subunit, which harbors the prosthetic group at its N terminus. The mechanism of PSD proenzyme cleavage has long been unclear. Using a coupled in vitro transcription/translation system with the soluble Plasmodium knowlesi enzyme (PkPSD), we demonstrate that the post-translational processing is inhibited by the serine protease inhibitor, phenylmethylsulfonyl fluoride. Comparison of PSD sequences across multiple phyla reveals a uniquely conserved aspartic acid within an FFXRX6RX12PXD motif, two uniquely conserved histidine residues within a PXXYHXXHXP motif, and a uniquely conserved serine residue within a GS(S/T) motif, suggesting that PSDs belong to the D-H-S serine protease family. The function of the conserved D-H-S residues was probed using site-directed mutagenesis of PkPSD. The results from these mutagenesis experiments reveal that Asp-139, His-198, and Ser-308 are all essential for endoproteolytic processing of PkPSD, which occurs in cis. In addition, within the GS(S/T) motif found in all PSDs, the Gly-307 residue is also essential, but the Ser/Thr-309 is non-essential. These results define the mechanism whereby PSDs begin their biochemical existence as proteases that execute one autoendoproteolytic cleavage reaction to give rise to a mature PSD harboring a pyruvoyl prosthetic group.
Keywords: Autoendoproteolysis; Membrane; Phosphatidylserine Decarboxylase; Phospholipid; Plasmodium; Protease; Pyruvoyl-enzyme.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures
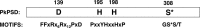



Similar articles
-
Maturation of the malarial phosphatidylserine decarboxylase is mediated by high affinity binding to anionic phospholipids.J Biol Chem. 2023 May;299(5):104659. doi: 10.1016/j.jbc.2023.104659. Epub 2023 Mar 29. J Biol Chem. 2023. PMID: 36997087 Free PMC article.
-
Identification of gene encoding Plasmodium knowlesi phosphatidylserine decarboxylase by genetic complementation in yeast and characterization of in vitro maturation of encoded enzyme.J Biol Chem. 2012 Jan 2;287(1):222-232. doi: 10.1074/jbc.M111.313676. Epub 2011 Nov 4. J Biol Chem. 2012. PMID: 22057268 Free PMC article.
-
High-throughput screening for phosphatidylserine decarboxylase inhibitors using a distyrylbenzene-bis-aldehyde (DSB-3)-based fluorescence assay.J Biol Chem. 2019 Aug 9;294(32):12146-12156. doi: 10.1074/jbc.RA119.007766. Epub 2019 Jun 21. J Biol Chem. 2019. PMID: 31227523 Free PMC article.
-
Phosphatidylserine decarboxylase.Biochim Biophys Acta. 1997 Sep 4;1348(1-2):236-44. doi: 10.1016/s0005-2760(97)00101-x. Biochim Biophys Acta. 1997. PMID: 9370338 Review.
-
Phosphatidylserine decarboxylases, key enzymes of lipid metabolism.IUBMB Life. 2009 Feb;61(2):151-62. doi: 10.1002/iub.159. IUBMB Life. 2009. PMID: 19165886 Review.
Cited by
-
Structural insights into phosphatidylethanolamine formation in bacterial membrane biogenesis.Sci Rep. 2021 Mar 11;11(1):5785. doi: 10.1038/s41598-021-85195-5. Sci Rep. 2021. PMID: 33707636 Free PMC article.
-
Preparation of Asymmetric Liposomes Using a Phosphatidylserine Decarboxylase.Biophys J. 2018 Oct 16;115(8):1509-1517. doi: 10.1016/j.bpj.2018.08.036. Epub 2018 Sep 1. Biophys J. 2018. PMID: 30266319 Free PMC article.
-
The catalytic and structural basis of archaeal glycerophospholipid biosynthesis.Extremophiles. 2022 Aug 17;26(3):29. doi: 10.1007/s00792-022-01277-w. Extremophiles. 2022. PMID: 35976526 Free PMC article. Review.
-
Structural basis for psilocybin biosynthesis.Nat Commun. 2025 Mar 22;16(1):2827. doi: 10.1038/s41467-025-58239-x. Nat Commun. 2025. PMID: 40121242 Free PMC article.
-
Interaction of two antitumor peptides with membrane lipids - Influence of phosphatidylserine and cholesterol on specificity for melanoma cells.PLoS One. 2019 Jan 25;14(1):e0211187. doi: 10.1371/journal.pone.0211187. eCollection 2019. PLoS One. 2019. PMID: 30682171 Free PMC article.
References
-
- Voelker D. R. (1997) Phosphatidylserine decarboxylase. Biochim. Biophys. Acta 1348, 236–244 - PubMed
-
- Kennedy E. P., Weiss S. B. (1956) The function of cytidine coenzymes in the biosynthesis of phospholipides. J. Biol. Chem. 222, 193–214 - PubMed
-
- Vance J. E., Tasseva G. (2013) Formation and function of phosphatidylserine and phosphatidylethanolamine in mammalian cells. Biochim. Biophys. Acta 1831, 543–554 - PubMed
-
- Sundler R., Akesson B., Nilsson A. (1974) Quantitative role of base exchange in phosphatidylethanolamine synthesis in isolated rat hepatocytes. FEBS Lett. 43, 303–307 - PubMed
-
- Riekhof W. R., Voelker D. R. (2006) Uptake and utilization of lyso-phosphatidylethanolamine by Saccharomyces cerevisiae. J. Biol. Chem. 281, 36588–36596 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

