Structure-activity study for (bis)ureidopropyl- and (bis)thioureidopropyldiamine LSD1 inhibitors with 3-5-3 and 3-6-3 carbon backbone architectures
- PMID: 25725609
- PMCID: PMC4396983
- DOI: 10.1016/j.bmc.2015.01.049
Structure-activity study for (bis)ureidopropyl- and (bis)thioureidopropyldiamine LSD1 inhibitors with 3-5-3 and 3-6-3 carbon backbone architectures
Erratum in
-
Corrigendum to "Structure-activity study for (bis)ureidopropyl- and (bis)thioureidopropyldiamine LSD1 inhibitors with 3-5-3 and 3-6-3 carbon backbone architectures" [Bioorg. Med. Chem. 23 (2015) 1601-1612].Bioorg Med Chem. 2018 Jul 23;26(12):3728-3729. doi: 10.1016/j.bmc.2018.05.032. Epub 2018 May 30. Bioorg Med Chem. 2018. PMID: 29859749 No abstract available.
Abstract
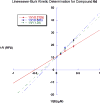
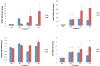
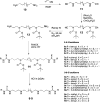
Methylation at specific histone lysine residues is a critical post-translational modification that alters chromatin architecture, and dysregulated lysine methylation/demethylation is associated with the silencing of tumor suppressor genes. The enzyme lysine-specific demethylase 1 (LSD1) complexed to specific transcription factors catalyzes the oxidative demethylation of mono- and dimethyllysine 4 of histone H3 (H3K4me and H3K4me2, respectively). We have previously reported potent (bis)urea and (bis)thiourea LSD1 inhibitors that increase cellular levels of H3K4me and H3K4me2, promote the re-expression of silenced tumor suppressor genes and suppress tumor growth in vitro. Here we report the design additional (bis)urea and (bis)thiourea LSD1 inhibitors that feature 3-5-3 or 3-6-3 carbon backbone architectures. Three of these compounds displayed single-digit IC50 values in a recombinant LSD1 assay. In addition, compound 6d exhibited an IC50 of 4.2μM against the Calu-6 human lung adenocarcinoma line, and 4.8μM against the MCF7 breast tumor cell line, in an MTS cell viability assay. Following treatment with 6b-6d, Calu-6 cells exhibited a significant increase in the mRNA expression for the silenced tumor suppressor genes SFRP2, HCAD and p16, and modest increases in GATA4 message. The compounds described in this paper represent the most potent epigenetic modulators in this series, and have potential for use as antitumor agents.
Keywords: Antitumor agent; Epigenetics; Histone demethylase; Lysine-specific demethylase 1; Oligoamine.
Copyright © 2015 Elsevier Ltd. All rights reserved.
Figures


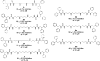


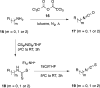
References
-
- Marks P, Rifkind RA, Richon VM, Breslow R, Miller T, Kelly WK. Nat Rev Cancer. 2001;1:194. - PubMed
-
- Shi Y, Whetstine JR. Molec Cell. 2007;25:1. - PubMed
-
- Herman JG, Baylin SB. N Engl J Med. 2003;349:2042. - PubMed
-
- Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA, Shi Y. Cell. 2004;119:941. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Chemical Information
Miscellaneous

