Gene Signatures of 1,25-Dihydroxyvitamin D3 Exposure in Normal and Transformed Mammary Cells
- PMID: 25736056
- PMCID: PMC6607438
- DOI: 10.1002/jcb.25129
Gene Signatures of 1,25-Dihydroxyvitamin D3 Exposure in Normal and Transformed Mammary Cells
Abstract
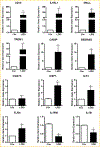
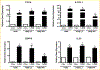
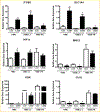
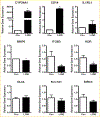
To elucidate potential mediators of vitamin D receptor (VDR) action in breast cancer, we profiled the genomic effects of its ligand 1,25-dihydroxyvitamin D3 (1,25D) in cells derived from normal mammary tissue and breast cancer. In non-transformed hTERT-HME cells, 483 1,25D responsive entities in 42 pathways were identified, whereas in MCF7 breast cancer cells, 249 1,25D responsive entities in 31 pathways were identified. Only 21 annotated genes were commonly altered by 1,25D in both MCF7 and hTERT-HME cells. Gene set enrichment analysis highlighted eight pathways (including senescence/autophagy, TGFβ signaling, endochondral ossification, and adipogenesis) commonly altered by 1,25D in hTERT-HME and MCF7 cells. Regulation of a subset of immune (CD14, IL1RL1, MALL, CAMP, SEMA6D, TREM1, CSF1, IL33, TLR4) and metabolic (ITGB3, SLC1A1, G6PD, GLUL, HIF1A, KDR, BIRC3) genes by 1,25D was confirmed in hTERT-HME cells and similar changes were observed in another comparable non-transformed mammary cell line (HME cells). The effects of 1,25D on these genes were retained in HME cells expressing SV40 large T antigen but were selectively abrogated in HME cells expressing SV40 + RAS and in MCF7 cells. Integration of the datasets from hTERT-HME and MCF7 cells with publically available RNA-SEQ data from 1,25D treated SKBR3 breast cancer cells enabled identification of an 11-gene signature representative of 1,25D exposure in all three breast-derived cell lines. Four of these 11 genes (CYP24A1, CLMN, EFTUD1, and SERPINB1) were also identified as 1,25D responsive in human breast tumor explants, suggesting that this gene signature may prove useful as a biomarker of vitamin D exposure in breast tissue.
Keywords: BREAST CANCER; GENOMIC PROFILING; GROWTH INHIBITION; MAMMARY EPITHELIAL CELLS; MICROARRAY; VITAMIN D RECEPTOR.
© 2015 Wiley Periodicals, Inc.
Figures


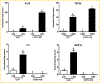


References
-
- Aoyama K, Suh SW, Hamby AM, Liu J, Chan WY, Chen Y, Swanson RA. 2006. Neuronal glutathione deficiency and age-dependent neurodegeneration in the EAAC1 deficient mouse. Nat Neurosci 9:119–126. - PubMed
-
- Beaudin SG, Robilotto S, Welsh J. 2014. Comparative regulation of gene expression by 1,25-dihydroxyvitamin D in cells derived from normal mammary tissue and breast cancer. J Steroid Biochem Mol Biol Sep 18. pii: S0960–0760(14) 00213–1 Sep 18. pii: S0960–0760(14) 00213–1. doi: 10.1016/j.jsbmb.2014.09.014. [Epub ahead of print]. - DOI - PMC - PubMed
-
- Bensaad K, Harris AL. 2014. Hypoxia and metabolism in cancer. Adv Exp Med Biol 772:1–39. - PubMed
-
- Berger U, Wilson P, McClelland RA, Colston K, Haussler MR, Pike JW, Coombes RC. 1987. Immunocytochemical detection of 1,25-dihydroxyvita-min D3 receptor in breast cancer. Cancer Res 47:6793–6799. - PubMed
-
- Buras RR, Schumaker LM, Davoodi F, Brenner RV, Shabahang M, Nauta RJ, Evans SR. 1994. Vitamin D receptors in breast cancer cells. Breast Cancer Res Treat 31:191–202. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

