Role of integrons, plasmids and SXT elements in multidrug resistance of Vibrio cholerae and Providencia vermicola obtained from a clinical isolate of diarrhea
- PMID: 25741322
- PMCID: PMC4330914
- DOI: 10.3389/fmicb.2015.00057
Role of integrons, plasmids and SXT elements in multidrug resistance of Vibrio cholerae and Providencia vermicola obtained from a clinical isolate of diarrhea
Abstract
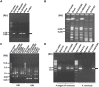
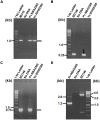
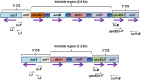
The isolates of Vibrio cholerae and Providencia vermicola obtained from a diarrheal patient were investigated for genetic elements governing their drug resistance phenotypes. Out of 14 antibiotics tested, V. cholerae Vc IDH02365 isolate showed resistance to nine antibiotics, while P. vermicola Pv NBA2365 was found to be resistant to all the antibiotics except polymyxin B. Though SXT integrase was depicted in both the bacteria, class 1 integron was found to be associated only with Pv NBA2365. Integrons in Pv NBA2365 conferred resistance to β-lactams, aminoglycosides, and trimethoprim. Pv NBA2365 carried two transformable plasmids imparting distinct antibiotic resistance traits to their Escherichia coli transformants. In rabbit ileal loop assays, Pv NBA2365 did not show any fluid accumulation (FA) in contrast with Vc IDH02365 that showed high FA. To the best of our knowledge, this is the first report of a highly drug resistant P. vermicola and additionally co-existence of multidrug resistant V. cholerae and P. vermicola. Both the microbes appeared to possess a wide array of mobile genetic elements for a large spectrum of antimicrobial agents, some of which are being used in the treatment of acute diarrhea.
Keywords: Providencia vermicola; SXT element; Vibrio cholerae; integron; multidrug resistance; plasmid.
Figures


Similar articles
-
New cluster of plasmid-located class 1 integrons in Vibrio cholerae O1 and a dfrA15 cassette-containing integron in Vibrio parahaemolyticus isolated in Angola.Antimicrob Agents Chemother. 2006 Jul;50(7):2493-9. doi: 10.1128/AAC.01310-05. Antimicrob Agents Chemother. 2006. PMID: 16801431 Free PMC article.
-
IncA/C Conjugative Plasmids Mobilize a New Family of Multidrug Resistance Islands in Clinical Vibrio cholerae Non-O1/Non-O139 Isolates from Haiti.mBio. 2016 Jul 19;7(4):e00509-16. doi: 10.1128/mBio.00509-16. mBio. 2016. PMID: 27435459 Free PMC article.
-
Mechanism of drug resistance in a clinical isolate of Vibrio fluvialis: involvement of multiple plasmids and integrons.Int J Antimicrob Agents. 2009 Sep;34(3):220-5. doi: 10.1016/j.ijantimicag.2009.03.020. Epub 2009 May 7. Int J Antimicrob Agents. 2009. PMID: 19427174
-
Antibiotic resistance mechanisms of Vibrio cholerae.J Med Microbiol. 2011 Apr;60(Pt 4):397-407. doi: 10.1099/jmm.0.023051-0. Epub 2011 Jan 20. J Med Microbiol. 2011. PMID: 21252269 Review.
-
Antibiotic Resistance Conferred by Class 1 Integron in Vibrio Cholerae Strains: A Meta-analysis.East Afr Health Res J. 2022;6(2):119-126. doi: 10.24248/eahrj.v6i2.690. Epub 2022 Nov 30. East Afr Health Res J. 2022. PMID: 36751685 Free PMC article. Review.
Cited by
-
Antibiotic resistance profiling of pathogenic Enterobacteriaceae from Cluj-Napoca, Romania.Germs. 2019 Mar 1;9(1):17-27. doi: 10.18683/germs.2019.1153. eCollection 2019 Mar. Germs. 2019. PMID: 31119113 Free PMC article.
-
Comparative Genomics Reveals High Genomic Diversity in the Genus Photobacterium.Front Microbiol. 2017 Jun 29;8:1204. doi: 10.3389/fmicb.2017.01204. eCollection 2017. Front Microbiol. 2017. PMID: 28706512 Free PMC article.
-
Isolation and Molecular Identification and Antimicrobial Susceptibility of Providencia spp. from Raw Cow's Milk in Baghdad, Iraq.Vet Med Int. 2020 Nov 19;2020:8874747. doi: 10.1155/2020/8874747. eCollection 2020. Vet Med Int. 2020. PMID: 33456748 Free PMC article.
-
First Genome Description of Providencia vermicola Isolate Bearing NDM-1 from Blood Culture.Microorganisms. 2021 Aug 17;9(8):1751. doi: 10.3390/microorganisms9081751. Microorganisms. 2021. PMID: 34442831 Free PMC article.
-
Genomic analysis of the nomenclatural type strain of the nematode-associated entomopathogenic bacterium Providencia vermicola.BMC Genomics. 2021 Oct 2;22(1):708. doi: 10.1186/s12864-021-08027-w. BMC Genomics. 2021. PMID: 34598677 Free PMC article.
References
-
- Atlas R. M. (2010). Handbook of Microbiological Media 4th Edn. (Boca Raton, FL: CRC Press Inc.).
-
- Bhardwaj A. K., Vinothkumar K. (2015). “Evolution of MDRs,” in Quorum Sensing vs Quorum Quenching: A Battle with No End in Sight ed. Kalia V. C. (New Delhi: Springer) 9–22 10.1007/978-81-322-1982-8_2 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

