Systems biology approaches for advancing the discovery of effective drug combinations
- PMID: 25741385
- PMCID: PMC4348553
- DOI: 10.1186/s13321-015-0055-9
Systems biology approaches for advancing the discovery of effective drug combinations
Abstract
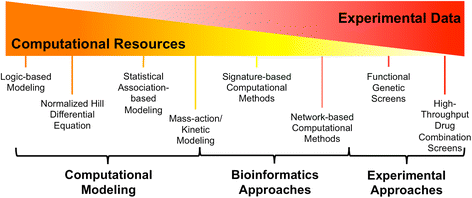
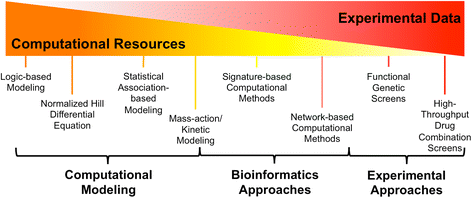
Complex diseases like cancer are regulated by large, interconnected networks with many pathways affecting cell proliferation, invasion, and drug resistance. However, current cancer therapy predominantly relies on the reductionist approach of one gene-one disease. Combinations of drugs may overcome drug resistance by limiting mutations and induction of escape pathways, but given the enormous number of possible drug combinations, strategies to reduce the search space and prioritize experiments are needed. In this review, we focus on the use of computational modeling, bioinformatics and high-throughput experimental methods for discovery of drug combinations. We highlight cutting-edge systems approaches, including large-scale modeling of cell signaling networks, network motif analysis, statistical association-based models, identifying correlations in gene signatures, functional genomics, and high-throughput combination screens. We also present a list of publicly available data and resources to aid in discovery of drug combinations. Integration of these systems approaches will enable faster discovery and translation of clinically relevant drug combinations. Graphical abstractSpectrum of Systems Biology Approaches for Drug Combinations.
Keywords: Cancer; Computational modeling; Drug combinations; Drug discovery; Systems biology.
Figures
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

