Genomics of injury: The Glue Grant experience
- PMID: 25742245
- PMCID: PMC4389222
- DOI: 10.1097/TA.0000000000000568
Genomics of injury: The Glue Grant experience
Figures



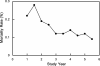



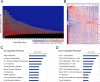
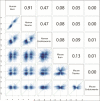
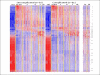
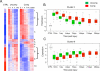

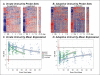
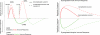
References
-
- Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA, et al. the Human Genome Project Investigators The sequence of the human genome. Science. 2001 Feb;291:1304–51. - PubMed
-
- Schena M, Shalon D, Davis RW, Brown PO. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995 Oct;270:467–70. - PubMed
-
- The International HapMap Consortium The international hapmap project. Nature. 2003 Dec;246:789–96. - PubMed
-
- Bone RC. A piece of my mind. Maumee: my Walden pond. JAMA. 1996 Dec;276:1931. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

