NAD(+)-SIRT1 control of H3K4 trimethylation through circadian deacetylation of MLL1
- PMID: 25751424
- PMCID: PMC4732879
- DOI: 10.1038/nsmb.2990
NAD(+)-SIRT1 control of H3K4 trimethylation through circadian deacetylation of MLL1
Abstract
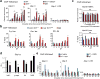
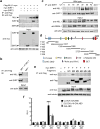
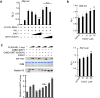
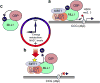
The circadian clock controls the transcription of hundreds of genes through specific chromatin-remodeling events. The histone methyltransferase mixed-lineage leukemia 1 (MLL1) coordinates recruitment of CLOCK-BMAL1 activator complexes to chromatin, an event associated with cyclic trimethylation of histone H3 Lys4 (H3K4) at circadian promoters. Remarkably, in mouse liver circadian H3K4 trimethylation is modulated by SIRT1, an NAD(+)-dependent deacetylase involved in clock control. We show that mammalian MLL1 is acetylated at two conserved residues, K1130 and K1133. Notably, MLL1 acetylation is cyclic, controlled by the clock and by SIRT1, and it affects the methyltransferase activity of MLL1. Moreover, H3K4 methylation at clock-controlled-gene promoters is influenced by pharmacological or genetic inactivation of SIRT1. Finally, levels of MLL1 acetylation and H3K4 trimethylation at circadian gene promoters depend on NAD(+) circadian levels. These findings reveal a previously unappreciated regulatory pathway between energy metabolism and histone methylation.
Figures

Comment in
-
Methylation gets into rhythm with NAD(+)-SIRT1.Nat Struct Mol Biol. 2015 Apr;22(4):275-7. doi: 10.1038/nsmb.3004. Nat Struct Mol Biol. 2015. PMID: 25837871
References
-
- Ripperger JA, Schibler U. Rhythmic CLOCK–BMAL1 binding to multiple E-box motifs drives circadian Dbp transcription and chromatin transitions. Nature genetics. 2006;38:369–74. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

