A putative cro-like repressor contributes to arylomycin resistance in Staphylococcus aureus
- PMID: 25753642
- PMCID: PMC4432125
- DOI: 10.1128/AAC.04597-14
A putative cro-like repressor contributes to arylomycin resistance in Staphylococcus aureus
Abstract
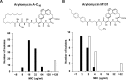
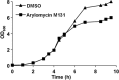
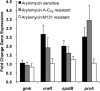
Antibiotic-resistant bacteria are a significant public health concern and motivate efforts to develop new classes of antibiotics. One such class of antibiotics is the arylomycins, which target type I signal peptidase (SPase), the enzyme responsible for the release of secreted proteins from their N-terminal leader sequences. Despite the essentiality, conservation, and relative accessibility of SPase, the activity of the arylomycins is limited against some bacteria, including the important human pathogen Staphylococcus aureus. To understand the origins of the limited activity against S. aureus, we characterized the susceptibility of a panel of strains to two arylomycin derivatives, arylomycin A-C16 and its more potent analog arylomycin M131. We observed a wide range of susceptibilities to the two arylomycins and found that resistant strains were sensitized by cotreatment with tunicamycin, which inhibits the first step of wall teichoic acid synthesis. To further understand how S. aureus responds to the arylomycins, we profiled the transcriptional response of S. aureus NCTC 8325 to growth-inhibitory concentrations of arylomycin M131 and found that it upregulates the cell wall stress stimulon (CWSS) and an operon consisting of a putative transcriptional regulator and three hypothetical proteins. Interestingly, we found that mutations in the putative transcriptional regulator are correlated with resistance, and selection for resistance ex vivo demonstrated that mutations in this gene are sufficient for resistance. The results begin to elucidate how S. aureus copes with secretion stress and how it evolves resistance to the inhibition of SPase.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
A Putative Bacterial ABC Transporter Circumvents the Essentiality of Signal Peptidase.mBio. 2016 Sep 6;7(5):e00412-16. doi: 10.1128/mBio.00412-16. mBio. 2016. PMID: 27601569 Free PMC article.
-
Type I signal peptidase and protein secretion in Staphylococcus aureus.J Bacteriol. 2012 May;194(10):2677-86. doi: 10.1128/JB.00064-12. Epub 2012 Mar 23. J Bacteriol. 2012. PMID: 22447899 Free PMC article.
-
Mechanism of action of the arylomycin antibiotics and effects of signal peptidase I inhibition.Antimicrob Agents Chemother. 2012 Oct;56(10):5054-60. doi: 10.1128/AAC.00785-12. Epub 2012 Jul 16. Antimicrob Agents Chemother. 2012. PMID: 22802255 Free PMC article.
-
Bacterial type I signal peptidase inhibitors - Optimized hits from nature.Eur J Med Chem. 2022 Aug 5;238:114490. doi: 10.1016/j.ejmech.2022.114490. Epub 2022 May 26. Eur J Med Chem. 2022. PMID: 35660251 Review.
-
Mechanisms of antibiotic resistance in Staphylococcus aureus.Future Microbiol. 2007 Jun;2(3):323-34. doi: 10.2217/17460913.2.3.323. Future Microbiol. 2007. PMID: 17661706 Review.
Cited by
-
The inhibition of type I bacterial signal peptidase: Biological consequences and therapeutic potential.Bioorg Med Chem Lett. 2015 Nov 1;25(21):4761-4766. doi: 10.1016/j.bmcl.2015.07.072. Epub 2015 Jul 26. Bioorg Med Chem Lett. 2015. PMID: 26276537 Free PMC article. Review.
-
Solid-Phase Synthesis of Biaryl Cyclic Lipopeptides Derived from Arylomycins.ACS Omega. 2020 Sep 4;5(36):23401-23412. doi: 10.1021/acsomega.0c03352. eCollection 2020 Sep 15. ACS Omega. 2020. PMID: 32954193 Free PMC article.
-
A Putative Bacterial ABC Transporter Circumvents the Essentiality of Signal Peptidase.mBio. 2016 Sep 6;7(5):e00412-16. doi: 10.1128/mBio.00412-16. mBio. 2016. PMID: 27601569 Free PMC article.
-
Staphylococcus aureus type I signal peptidase: essential or not essential, that's the question.Microb Cell. 2017 Mar 17;4(4):108-111. doi: 10.15698/mic2017.04.566. Microb Cell. 2017. PMID: 28435837 Free PMC article.
-
A Review of Antibacterial Candidates with New Modes of Action.ACS Infect Dis. 2024 Oct 11;10(10):3440-3474. doi: 10.1021/acsinfecdis.4c00218. Epub 2024 Jul 17. ACS Infect Dis. 2024. PMID: 39018341 Free PMC article. Review.
References
-
- Hancock REW. 2007. The end of an era? Nat Rev Drug Discov 6:28. doi:10.1038/nrd2223. - DOI
-
- Wooldridge K. (ed). 2009. Bacterial secreted proteins: secretory mechanisms and role in pathogenesis. Caister Academic Press, Norwich, United Kingdom.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

