Improved cell-penetrating zinc-finger nuclease proteins for precision genome engineering
- PMID: 25756962
- PMCID: PMC4354341
- DOI: 10.1038/mtna.2015.6
Improved cell-penetrating zinc-finger nuclease proteins for precision genome engineering
Abstract
Safe, efficient, and broadly applicable methods for delivering site-specific nucleases into cells are needed in order for targeted genome editing to reach its full potential for basic research and medicine. We previously reported that zinc-finger nuclease (ZFN) proteins have the innate capacity to cross cell membranes and induce genome modification via their direct application to human cells. Here, we show that incorporation of tandem nuclear localization signal (NLS) repeats into the ZFN protein backbone enhances cell permeability nearly 13-fold and that single administration of multi-NLS ZFN proteins leads to genome modification rates of up to 26% in CD4(+) T cells and 17% in CD34(+) hematopoietic stem/progenitor cells. In addition, we show that multi-NLS ZFN proteins attenuate off-target effects and that codelivery of ZFN protein pairs facilitates dual gene modification frequencies of 20-30% in CD4(+) T cells. These results illustrate the applicability of ZFN protein delivery for precision genome engineering.
Figures


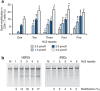
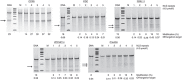
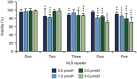

References
-
- Segal DJ, Meckler JF. Genome engineering at the dawn of the golden age. Annu Rev Genomics Hum Genet. 2013;14:135–158. - PubMed
-
- Kim H, Kim JS. A guide to genome engineering with programmable nucleases. Nat Rev Genet. 2014;15:321–334. - PubMed
-
- Pâques F, Duchateau P. Meganucleases and DNA double-strand break-induced recombination: perspectives for gene therapy. Curr Gene Ther. 2007;7:49–66. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

