Transcriptome analysis of thermophilic methylotrophic Bacillus methanolicus MGA3 using RNA-sequencing provides detailed insights into its previously uncharted transcriptional landscape
- PMID: 25758049
- PMCID: PMC4342826
- DOI: 10.1186/s12864-015-1239-4
Transcriptome analysis of thermophilic methylotrophic Bacillus methanolicus MGA3 using RNA-sequencing provides detailed insights into its previously uncharted transcriptional landscape
Abstract
Background: Bacillus methanolicus MGA3 is a thermophilic, facultative ribulose monophosphate (RuMP) cycle methylotroph. Together with its ability to produce high yields of amino acids, the relevance of this microorganism as a promising candidate for biotechnological applications is evident. The B. methanolicus MGA3 genome consists of a 3,337,035 nucleotides (nt) circular chromosome, the 19,174 nt plasmid pBM19 and the 68,999 nt plasmid pBM69. 3,218 protein-coding regions were annotated on the chromosome, 22 on pBM19 and 82 on pBM69. In the present study, the RNA-seq approach was used to comprehensively investigate the transcriptome of B. methanolicus MGA3 in order to improve the genome annotation, identify novel transcripts, analyze conserved sequence motifs involved in gene expression and reveal operon structures. For this aim, two different cDNA library preparation methods were applied: one which allows characterization of the whole transcriptome and another which includes enrichment of primary transcript 5'-ends.
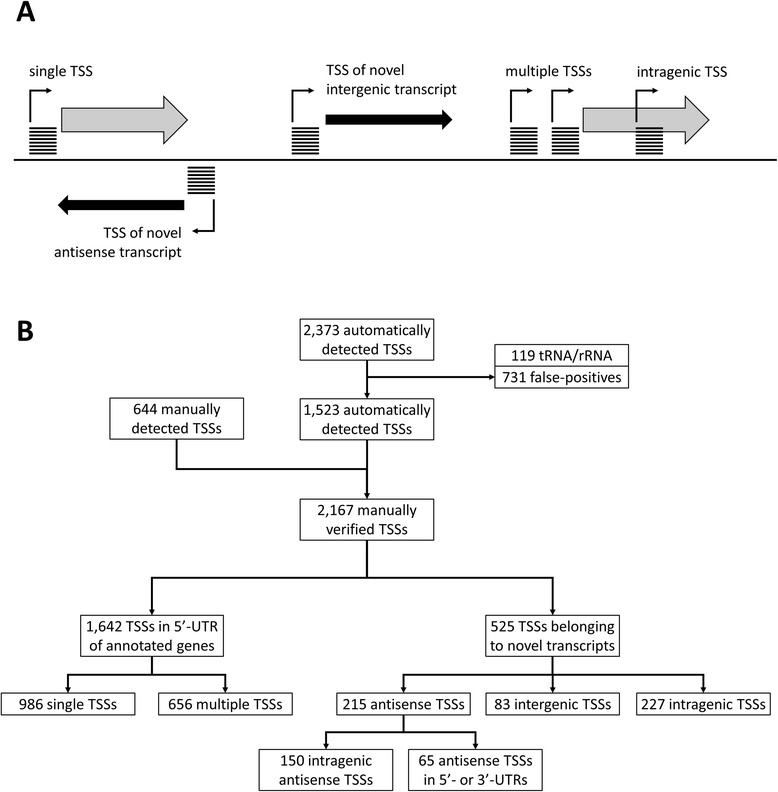
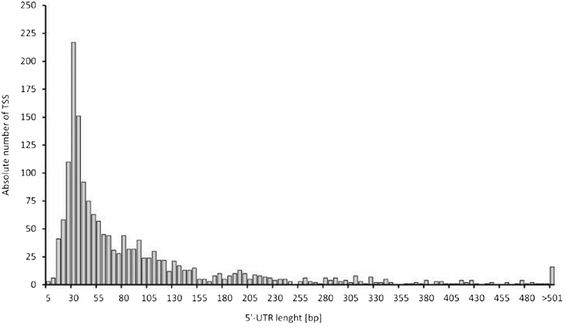
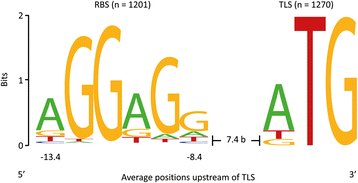
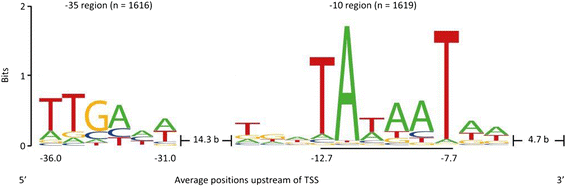
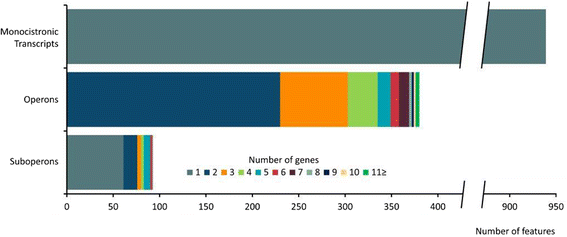
Results: Analysis of the primary transcriptome data enabled the detection of 2,167 putative transcription start sites (TSSs) which were categorized into 1,642 TSSs located in the upstream region (5'-UTR) of known protein-coding genes and 525 TSSs of novel antisense, intragenic, or intergenic transcripts. Firstly, 14 wrongly annotated translation start sites (TLSs) were corrected based on primary transcriptome data. Further investigation of the identified 5'-UTRs resulted in the detailed characterization of their length distribution and the detection of 75 hitherto unknown cis-regulatory RNA elements. Moreover, the exact TSSs positions were utilized to define conserved sequence motifs for translation start sites, ribosome binding sites and promoters in B. methanolicus MGA3. Based on the whole transcriptome data set, novel transcripts, operon structures and mRNA abundances were determined. The analysis of the operon structures revealed that almost half of the genes are transcribed monocistronically (940), whereas 1,164 genes are organized in 381 operons. Several of the genes related to methylotrophy had highly abundant transcripts.
Conclusion: The extensive insights into the transcriptional landscape of B. methanolicus MGA3, gained in this study, represent a valuable foundation for further comparative quantitative transcriptome analyses and possibly also for the development of molecular biology tools which at present are very limited for this organism.
Figures
Similar articles
-
Complete genome sequence of Bacillus methanolicus MGA3, a thermotolerant amino acid producing methylotroph.J Biotechnol. 2014 Oct 20;188:110-1. doi: 10.1016/j.jbiotec.2014.08.013. Epub 2014 Aug 22. J Biotechnol. 2014. PMID: 25152427
-
6-Phosphofructokinase and ribulose-5-phosphate 3-epimerase in methylotrophic Bacillus methanolicus ribulose monophosphate cycle.Appl Microbiol Biotechnol. 2017 May;101(10):4185-4200. doi: 10.1007/s00253-017-8173-0. Epub 2017 Feb 17. Appl Microbiol Biotechnol. 2017. PMID: 28213736
-
RNAseq analysis of α-proteobacterium Gluconobacter oxydans 621H.BMC Genomics. 2018 Jan 6;19(1):24. doi: 10.1186/s12864-017-4415-x. BMC Genomics. 2018. PMID: 29304737 Free PMC article.
-
Methylotrophy in the thermophilic Bacillus methanolicus, basic insights and application for commodity production from methanol.Appl Microbiol Biotechnol. 2015 Jan;99(2):535-51. doi: 10.1007/s00253-014-6224-3. Epub 2014 Nov 28. Appl Microbiol Biotechnol. 2015. PMID: 25431011 Review.
-
Bacillus methanolicus: an emerging chassis for low-carbon biomanufacturing.Trends Biotechnol. 2025 Feb;43(2):274-277. doi: 10.1016/j.tibtech.2024.06.013. Epub 2024 Jul 19. Trends Biotechnol. 2025. PMID: 39033040 Review.
Cited by
-
Extensive Reannotation of the Genome of the Model Streptomycete Streptomyces lividans TK24 Based on Transcriptome and Proteome Information.Front Microbiol. 2021 Apr 14;12:604034. doi: 10.3389/fmicb.2021.604034. eCollection 2021. Front Microbiol. 2021. PMID: 33935985 Free PMC article.
-
Transcriptome sequencing of the human pathogen Corynebacterium diphtheriae NCTC 13129 provides detailed insights into its transcriptional landscape and into DtxR-mediated transcriptional regulation.BMC Genomics. 2018 Jan 25;19(1):82. doi: 10.1186/s12864-018-4481-8. BMC Genomics. 2018. PMID: 29370758 Free PMC article.
-
Structure of a Core Promoter in Bifidobacterium longum NCC2705.J Bacteriol. 2020 Mar 11;202(7):e00540-19. doi: 10.1128/JB.00540-19. Print 2020 Mar 11. J Bacteriol. 2020. PMID: 31964699 Free PMC article.
-
Time-resolved transcriptome analysis and lipid pathway reconstruction of the oleaginous green microalga Monoraphidium neglectum reveal a model for triacylglycerol and lipid hyperaccumulation.Biotechnol Biofuels. 2017 Aug 14;10:197. doi: 10.1186/s13068-017-0882-1. eCollection 2017. Biotechnol Biofuels. 2017. PMID: 28814974 Free PMC article.
-
Quantitative metabolomics of the thermophilic methylotroph Bacillus methanolicus.Microb Cell Fact. 2016 Jun 1;15:92. doi: 10.1186/s12934-016-0483-x. Microb Cell Fact. 2016. PMID: 27251037 Free PMC article.
References
-
- Arfman N, Dijkhuizen L, Kirchhof G, Ludwig W, Schleifer K, Bulygina ES, et al. Bacillus methanolicus sp. nov., a new species of thermotolerant, methanol-utilizing, endospore-forming bacteria. Int J Syst Bacteriol. 1992;42:439–45. - PubMed
-
- Schendel F, Dillingham R, Hanson RS, Sano K, Matsui K. Production of glutamate using wild type Bacillus methanolicus. Patent (US6083728). 2000:1–20
-
- Schrader J, Schilling M, Holtmann D, Sell D, Filho MV, Marx A, et al. Methanol-based industrial biotechnology: current status and future perspectives of methylotrophic bacteria. Trends Biotechnol. 2009;27:107–15. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

