Molecular biology of hepatitis B virus infection
- PMID: 25759099
- PMCID: PMC4424072
- DOI: 10.1016/j.virol.2015.02.031
Molecular biology of hepatitis B virus infection
Abstract
Human hepatitis B virus (HBV) is the prototype of a family of small DNA viruses that productively infect hepatocytes, the major cell of the liver, and replicate by reverse transcription of a terminally redundant viral RNA, the pregenome. Upon infection, the circular, partially double-stranded virion DNA is converted in the nucleus to a covalently closed circular DNA (cccDNA) that assembles into a minichromosome, the template for viral mRNA synthesis. Infection of hepatocytes is non-cytopathic. Infection of the liver may be either transient (<6 months) or chronic and lifelong, depending on the ability of the host immune response to clear the infection. Chronic infections can cause immune-mediated liver damage progressing to cirrhosis and hepatocellular carcinoma (HCC). The mechanisms of carcinogenesis are unclear. Antiviral therapies with nucleoside analog inhibitors of viral DNA synthesis delay sequelae, but cannot cure HBV infections due to the persistence of cccDNA in hepatocytes.
Keywords: Hepatitis B virus; Pathogenesis; Replication.
Copyright © 2015 Elsevier Inc. All rights reserved.
Figures

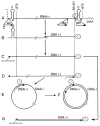



References
-
- Afdhal N, Zeuzem S, Kwo P, Chojkier M, Gitlin N, Puoti M, Romero-Gomez M, Zarski JP, Agarwal K, Buggisch P, Foster GR, Brau N, Buti M, Jacobson IM, Subramanian GM, Ding X, Mo H, Yang JC, Pang PS, Symonds WT, McHutchison JG, Muir AJ, Mangia A, Marcellin P. Ledipasvir and sofosbuvir for untreated HCV genotype 1 infection. NEJM. 2014;370:1889–1898. - PubMed
-
- Alter HJ, Holland PV, Purcell RH, Lander JJ, Feinstone SM, Morrow AG, Schmidt PJ. Posttransfusion hepatitis after exclusion of commercial and hepatitis-B antigen-positive donors. Ann Int Med. 1972;77:691–699. - PubMed
-
- Bancroft WH, Snitbhan R, Scott RM, Tingpalapong M, Watson WT, Tanticharoenyos P, Karwacki JJ, Srimarut S. Transmission of hepatitis B virus to gibbons by exposure to human saliva containing hepatitis B surface antigen. J Inf Dis. 1977;135:79–85. - PubMed
-
- Barker LF, Chisari FV, McGrath PP, Dalgard DW, Kirschstein RL, Almeida JD, Edgington TS, Sharp DG, Peterson MR. Transmission of type B viral hepatitis to chimpanzees. J Infect Dis. 1973;127:648–652. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

