Detection of a novel, integrative aging process suggests complex physiological integration
- PMID: 25761112
- PMCID: PMC4356614
- DOI: 10.1371/journal.pone.0116489
Detection of a novel, integrative aging process suggests complex physiological integration
Abstract
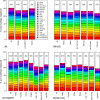
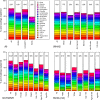
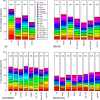
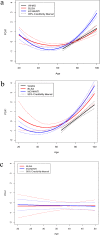
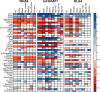
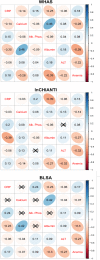
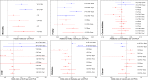
Many studies of aging examine biomarkers one at a time, but complex systems theory and network theory suggest that interpretations of individual markers may be context-dependent. Here, we attempted to detect underlying processes governing the levels of many biomarkers simultaneously by applying principal components analysis to 43 common clinical biomarkers measured longitudinally in 3694 humans from three longitudinal cohort studies on two continents (Women's Health and Aging I & II, InCHIANTI, and the Baltimore Longitudinal Study on Aging). The first axis was associated with anemia, inflammation, and low levels of calcium and albumin. The axis structure was precisely reproduced in all three populations and in all demographic sub-populations (by sex, race, etc.); we call the process represented by the axis "integrated albunemia." Integrated albunemia increases and accelerates with age in all populations, and predicts mortality and frailty--but not chronic disease--even after controlling for age. This suggests a role in the aging process, though causality is not yet clear. Integrated albunemia behaves more stably across populations than its component biomarkers, and thus appears to represent a higher-order physiological process emerging from the structure of underlying regulatory networks. If this is correct, detection of this process has substantial implications for physiological organization more generally.
Conflict of interest statement
Figures




References
-
- Harman SM, Metter EJ, Tobin JD, Pearson J, Blackman MR (2001) Longitudinal Effects of Aging on Serum Total and Free Testosterone Levels in Healthy Men. Journal of Clinical Endocrinology & Metabolism 86: 724–731. - PubMed
-
- Johnson TE (2006) Recent results: Biomarkers of aging. Experimental Gerontology 41: 1243–1246. - PubMed
-
- Finch CE (2007) The Biology of Human Longevity: Inflammation, Nutrition, and Aging in the Evolution of Life. San Diego, CA: Academic Press.
-
- Franceschi C, Bonafé M, Valensin S, Olivieri F, De Luca M, et al. (2000) Inflamm-aging: An Evolutionary Perspective on Immunosenescence. Annals of the New York Academy of Sciences 908: 244–254. - PubMed
-
- Cohen AA, Martin LB, Wingfield JC, McWilliams SR, Dunne JA (2012) Physiological regulatory networks: ecological roles and evolutionary constraints. Trends in Ecology & Evolution 27: 428–435. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

