Advanced fluorescence microscopy methods for the real-time study of transcription and chromatin dynamics
- PMID: 25764219
- PMCID: PMC4214231
- DOI: 10.4161/trns.28425
Advanced fluorescence microscopy methods for the real-time study of transcription and chromatin dynamics
Abstract
In this contribution we provide an overview of the recent advances allowed by the use of fluorescence microscopy methods in the study of transcriptional processes and their interplay with the chromatin architecture in living cells. Although the use of fluorophores to label nucleic acids dates back at least to about half a century ago, (1) two recent breakthroughs have effectively opened the way to use fluorescence routinely for specific and quantitative probing of chromatin organization and transcriptional activity in living cells: namely, the possibility of labeling first the chromatin loci and then the mRNA synthesized from a gene using fluorescent proteins. In this contribution we focus on methods that can probe rapid dynamic processes by analyzing fast fluorescence fluctuations.
Keywords: 3D orbital tracking; Fluorescence; Pair Correlation Function; RICS; brightness analysis; fluctuations.
Figures

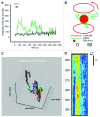
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
