Identification of U2AF(35)-dependent exons by RNA-Seq reveals a link between 3' splice-site organization and activity of U2AF-related proteins
- PMID: 25779042
- PMCID: PMC4402522
- DOI: 10.1093/nar/gkv194
Identification of U2AF(35)-dependent exons by RNA-Seq reveals a link between 3' splice-site organization and activity of U2AF-related proteins
Abstract
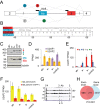
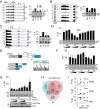
The auxiliary factor of U2 small nuclear RNA (U2AF) is a heterodimer consisting of 65- and 35-kD proteins that bind the polypyrimidine tract (PPT) and AG dinucleotides at the 3' splice site (3'ss). The gene encoding U2AF35 (U2AF1) is alternatively spliced, giving rise to two isoforms U2AF35a and U2AF35b. Here, we knocked down U2AF35 and each isoform and characterized transcriptomes of HEK293 cells with varying U2AF35/U2AF65 and U2AF35a/b ratios. Depletion of both isoforms preferentially modified alternative RNA processing events without widespread failure to recognize 3'ss or constitutive exons. Over a third of differentially used exons were terminal, resulting largely from the use of known alternative polyadenylation (APA) sites. Intronic APA sites activated in depleted cultures were mostly proximal whereas tandem 3'UTR APA was biased toward distal sites. Exons upregulated in depleted cells were preceded by longer AG exclusion zones and PPTs than downregulated or control exons and were largely activated by PUF60 and repressed by CAPERα. The U2AF(35) repression and activation was associated with a significant interchange in the average probabilities to form single-stranded RNA in the optimal PPT and branch site locations and sequences further upstream. Although most differentially used exons were responsive to both U2AF subunits and their inclusion correlated with U2AF levels, a small number of transcripts exhibited distinct responses to U2AF35a and U2AF35b, supporting the existence of isoform-specific interactions. These results provide new insights into function of U2AF and U2AF35 in alternative RNA processing.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




References
-
- Wahl M.C., Will C.L., Lührmann R. The spliceosome: design principles of a dynamic RNP machine. Cell. 2009;136:701–718. - PubMed
-
- Ruskin B., Zamore P.D., Green M.R. A factor, U2AF, is required for U2 snRNP binding and splicing complex assembly. Cell. 1988;52:207–219. - PubMed
-
- Zorio D.A., Blumenthal T. Both subunits of U2AF recognize the 3′ splice site in Caenorhabditis elegans. Nature. 1999;402:835–838. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

