Quantitative analysis of single-molecule force spectroscopy on folded chromatin fibers
- PMID: 25779043
- PMCID: PMC4402534
- DOI: 10.1093/nar/gkv215
Quantitative analysis of single-molecule force spectroscopy on folded chromatin fibers
Abstract
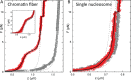
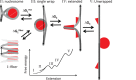
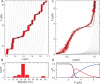
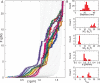
Single-molecule techniques allow for picoNewton manipulation and nanometer accuracy measurements of single chromatin fibers. However, the complexity of the data, the heterogeneity of the composition of individual fibers and the relatively large fluctuations in extension of the fibers complicate a structural interpretation of such force-extension curves. Here we introduce a statistical mechanics model that quantitatively describes the extension of individual fibers in response to force on a per nucleosome basis. Four nucleosome conformations can be distinguished when pulling a chromatin fiber apart. A novel, transient conformation is introduced that coexists with single wrapped nucleosomes between 3 and 7 pN. Comparison of force-extension curves between single nucleosomes and chromatin fibers shows that embedding nucleosomes in a fiber stabilizes the nucleosome by 10 kBT. Chromatin fibers with 20- and 50-bp linker DNA follow a different unfolding pathway. These results have implications for accessibility of DNA in fully folded and partially unwrapped chromatin fibers and are vital for understanding force unfolding experiments on nucleosome arrays.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
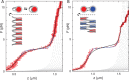
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

