Full-genome evolutionary histories of selfing, splitting, and selection in Caenorhabditis
- PMID: 25783854
- PMCID: PMC4417115
- DOI: 10.1101/gr.187237.114
Full-genome evolutionary histories of selfing, splitting, and selection in Caenorhabditis
Abstract
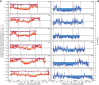
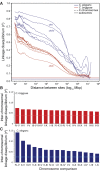
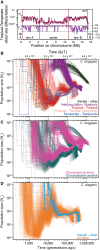
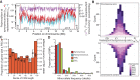
The nematode Caenorhabditis briggsae is a model for comparative developmental evolution with C. elegans. Worldwide collections of C. briggsae have implicated an intriguing history of divergence among genetic groups separated by latitude, or by restricted geography, that is being exploited to dissect the genetic basis to adaptive evolution and reproductive incompatibility; yet, the genomic scope and timing of population divergence is unclear. We performed high-coverage whole-genome sequencing of 37 wild isolates of the nematode C. briggsae and applied a pairwise sequentially Markovian coalescent (PSMC) model to 703 combinations of genomic haplotypes to draw inferences about population history, the genomic scope of natural selection, and to compare with 40 wild isolates of C. elegans. We estimate that a diaspora of at least six distinct C. briggsae lineages separated from one another approximately 200,000 generations ago, including the "Temperate" and "Tropical" phylogeographic groups that dominate most samples worldwide. Moreover, an ancient population split in its history approximately 2 million generations ago, coupled with only rare gene flow among lineage groups, validates this system as a model for incipient speciation. Low versus high recombination regions of the genome give distinct signatures of population size change through time, indicative of widespread effects of selection on highly linked portions of the genome owing to extreme inbreeding by self-fertilization. Analysis of functional mutations indicates that genomic context, owing to selection that acts on long linkage blocks, is a more important driver of population variation than are the functional attributes of the individually encoded genes.
© 2015 Thomas et al.; Published by Cold Spring Harbor Laboratory Press.
Figures


Similar articles
-
On the potential for extinction by Muller's ratchet in Caenorhabditis elegans.BMC Evol Biol. 2008 Apr 30;8:125. doi: 10.1186/1471-2148-8-125. BMC Evol Biol. 2008. PMID: 18447910 Free PMC article.
-
Selection at linked sites in the partial selfer Caenorhabditis elegans.Mol Biol Evol. 2003 May;20(5):665-73. doi: 10.1093/molbev/msg072. Epub 2003 Apr 2. Mol Biol Evol. 2003. PMID: 12679551
-
Natural selection shapes nucleotide polymorphism across the genome of the nematode Caenorhabditis briggsae.Genome Res. 2010 Aug;20(8):1103-11. doi: 10.1101/gr.104331.109. Epub 2010 May 27. Genome Res. 2010. PMID: 20508143 Free PMC article.
-
Molecular evolution inferences from the C. elegans genome.WormBook. 2010 Mar 5:1-14. doi: 10.1895/wormbook.1.149.1. WormBook. 2010. PMID: 20213859 Review.
-
Comparative genomics in C. elegans, C. briggsae, and other Caenorhabditis species.Methods Mol Biol. 2006;351:13-29. doi: 10.1385/1-59745-151-7:13. Methods Mol Biol. 2006. PMID: 16988423 Review.
Cited by
-
Domestication history and geographical adaptation inferred from a SNP map of African rice.Nat Genet. 2016 Sep;48(9):1083-8. doi: 10.1038/ng.3633. Epub 2016 Aug 8. Nat Genet. 2016. PMID: 27500524
-
Balancing selection maintains hyper-divergent haplotypes in Caenorhabditis elegans.Nat Ecol Evol. 2021 Jun;5(6):794-807. doi: 10.1038/s41559-021-01435-x. Epub 2021 Apr 5. Nat Ecol Evol. 2021. PMID: 33820969 Free PMC article.
-
Deep sampling of Hawaiian Caenorhabditis elegans reveals high genetic diversity and admixture with global populations.Elife. 2019 Dec 3;8:e50465. doi: 10.7554/eLife.50465. Elife. 2019. PMID: 31793880 Free PMC article.
-
Cold Survival and Its Molecular Mechanisms in a Locally Adapted Nematode Population.Genome Biol Evol. 2021 Sep 1;13(9):evab188. doi: 10.1093/gbe/evab188. Genome Biol Evol. 2021. PMID: 34383891 Free PMC article.
-
Divergent C. elegans toxin alleles are suppressed by distinct mechanisms.bioRxiv [Preprint]. 2024 Nov 20:2024.04.26.591160. doi: 10.1101/2024.04.26.591160. bioRxiv. 2024. PMID: 39605437 Free PMC article. Preprint.
References
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
