Self-assembly of a nine-residue amyloid-forming peptide fragment of SARS corona virus E-protein: mechanism of self aggregation and amyloid-inhibition of hIAPP
- PMID: 25785896
- PMCID: PMC4903029
- DOI: 10.1021/acs.biochem.5b00061
Self-assembly of a nine-residue amyloid-forming peptide fragment of SARS corona virus E-protein: mechanism of self aggregation and amyloid-inhibition of hIAPP
Abstract
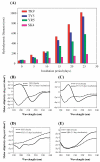
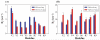
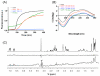
Molecular self-assembly, a phenomenon widely observed in nature, has been exploited through organic molecules, proteins, DNA, and peptides to study complex biological systems. These self-assembly systems may also be used in understanding the molecular and structural biology which can inspire the design and synthesis of increasingly complex biomaterials. Specifically, use of these building blocks to investigate protein folding and misfolding has been of particular value since it can provide tremendous insights into peptide aggregation related to a variety of protein misfolding diseases, or amyloid diseases (e.g., Alzheimer's disease, Parkinson's disease, type-II diabetes). Herein, the self-assembly of TK9, a nine-residue peptide of the extra membrane C-terminal tail of the SARS corona virus envelope, and its variants were characterized through biophysical, spectroscopic, and simulated studies, and it was confirmed that the structure of these peptides influences their aggregation propensity, hence, mimicking amyloid proteins. TK9, which forms a beta-sheet rich fibril, contains a key sequence motif that may be critical for beta-sheet formation, thus making it an interesting system to study amyloid fibrillation. TK9 aggregates were further examined through simulations to evaluate the possible intra- and interpeptide interactions at the molecular level. These self-assembly peptides can also serve as amyloid inhibitors through hydrophobic and electrophilic recognition interactions. Our results show that TK9 inhibits the fibrillation of hIAPP, a 37 amino acid peptide implicated in the pathology of type-II diabetes. Thus, biophysical and NMR experimental results have revealed a molecular level understanding of peptide folding events, as well as the inhibition of amyloid-protein aggregation are reported.
Figures



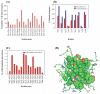
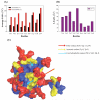
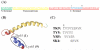
References
-
- Gazit E. Self-assembled peptide nanostructures: the design of molecular building blocks and their technological utilization. Chem. Soc. Rev. 2007;36:1263–1269. - PubMed
-
- Zhao X, Pan F, Xu H, Yaseen M, Shan H, Hauser CA, Zhang S, Lu JR. Molecular self-assembly and applications of designer peptide amphiphiles. Chem. Soc. Rev. 2010;39:3480–3498. - PubMed
-
- Zhang S. Fabrication of novel biomaterials through molecular self-assembly. Nat. Biotechnol. 2003;21:1171–1178. - PubMed
-
- Whitesides G, Mathias J, Seto C. Molecular self-assembly and nanochemistry: a chemical strategy for the synthesis of nanostructures. Science. 1991;254:1312–1319. - PubMed
-
- Lecommandoux S, Achard M-F, Langenwalter JF, Klok H-A. Self-assembly of rod-coil diblock oligomers based on α-helical peptides. Macromolecules. 2001;34:9100–9111.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

