Mechanochemical regulations of RPA's binding to ssDNA
- PMID: 25787788
- PMCID: PMC4365408
- DOI: 10.1038/srep09296
Mechanochemical regulations of RPA's binding to ssDNA
Abstract
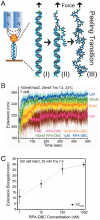
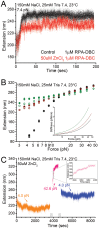
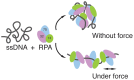
Replication protein A (RPA) is a ubiquitous eukaryotic single-stranded DNA (ssDNA) binding protein that serves to protect ssDNA from degradation and annealing, and as a template for recruitment of many downstream factors in virtually all DNA transactions in cell. During many of these transactions, DNA is tethered and is likely subject to force. Previous studies of RPA's binding behavior on ssDNA were conducted in the absence of force; therefore the RPA-ssDNA conformations regulated by force remain unclear. Here, using a combination of atomic force microscopy imaging and mechanical manipulation of single ssDNA tethers, we show that force mediates a switch of the RPA bound ssDNA from amorphous aggregation to a much more regular extended conformation. Further, we found an interesting non-monotonic dependence of the binding affinity on monovalent salt concentration in the presence of force. In addition, we discovered that zinc in micromolar concentrations drives ssDNA to a unique, highly stiff and more compact state. These results provide new mechanochemical insights into the influences and the mechanisms of action of RPA on large single ssDNA.
Figures


Similar articles
-
Role of zinc-finger motif in redox regulation of human replication protein A.Antioxid Redox Signal. 2001 Aug;3(4):657-69. doi: 10.1089/15230860152543005. Antioxid Redox Signal. 2001. PMID: 11554452
-
Phosphorylation of replication protein A by S-phase checkpoint kinases.DNA Repair (Amst). 2006 Mar 7;5(3):369-80. doi: 10.1016/j.dnarep.2005.11.007. Epub 2006 Jan 18. DNA Repair (Amst). 2006. PMID: 16412704
-
[Replication protein A as a major eukaryotic single-stranded DNA-binding protein and its role in DNA repair].Mol Biol (Mosk). 2016 Sep-Oct;50(5):735-750. doi: 10.7868/S0026898416030083. Mol Biol (Mosk). 2016. PMID: 27830676 Review. Russian.
-
A new structural framework for integrating replication protein A into DNA processing machinery.Nucleic Acids Res. 2013 Feb 1;41(4):2313-27. doi: 10.1093/nar/gks1332. Epub 2013 Jan 8. Nucleic Acids Res. 2013. PMID: 23303776 Free PMC article.
-
Replication Protein A, the Main Eukaryotic Single-Stranded DNA Binding Protein, a Focal Point in Cellular DNA Metabolism.Int J Mol Sci. 2024 Jan 2;25(1):588. doi: 10.3390/ijms25010588. Int J Mol Sci. 2024. PMID: 38203759 Free PMC article. Review.
Cited by
-
Human replication protein A induces dynamic changes in single-stranded DNA and RNA structures.J Biol Chem. 2019 Sep 20;294(38):13915-13927. doi: 10.1074/jbc.RA119.009737. Epub 2019 Jul 26. J Biol Chem. 2019. PMID: 31350334 Free PMC article.
-
Reduced Levels of Lagging Strand Polymerases Shape Stem Cell Chromatin.bioRxiv [Preprint]. 2024 Apr 29:2024.04.26.591383. doi: 10.1101/2024.04.26.591383. bioRxiv. 2024. Update in: Sci Adv. 2025 Feb 28;11(9):eadu6799. doi: 10.1126/sciadv.adu6799. PMID: 38746451 Free PMC article. Updated. Preprint.
-
Stabilization of DNA fork junctions by Smc5/6 complexes revealed by single-molecule imaging.Cell Rep. 2022 Dec 6;41(10):111778. doi: 10.1016/j.celrep.2022.111778. Cell Rep. 2022. PMID: 36476856 Free PMC article.
-
Oleandrin induces DNA damage responses in cancer cells by suppressing the expression of Rad51.Oncotarget. 2016 Sep 13;7(37):59572-59579. doi: 10.18632/oncotarget.10726. Oncotarget. 2016. PMID: 27449097 Free PMC article.
-
Parallel triplex structure formed between stretched single-stranded DNA and homologous duplex DNA.Nucleic Acids Res. 2017 Sep 29;45(17):10032-10041. doi: 10.1093/nar/gkx628. Nucleic Acids Res. 2017. PMID: 28973442 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

