Nuclear-encoded factors involved in post-transcriptional processing and modification of mitochondrial tRNAs in human disease
- PMID: 25806043
- PMCID: PMC4354410
- DOI: 10.3389/fgene.2015.00079
Nuclear-encoded factors involved in post-transcriptional processing and modification of mitochondrial tRNAs in human disease
Abstract
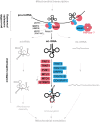
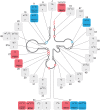
The human mitochondrial genome (mtDNA) encodes 22 tRNAs (mt-tRNAs) that are necessary for the intraorganellar translation of the 13 mtDNA-encoded subunits of the mitochondrial respiratory chain complexes. Maturation of mt-tRNAs involves 5' and 3' nucleolytic excision from precursor RNAs, as well as extensive post-transcriptional modifications. Recent data suggest that over 7% of all mt-tRNA residues in mammals undergo post-transcriptional modification, with over 30 different modified mt-tRNA positions so far described. These processing and modification steps are necessary for proper mt-tRNA function, and are performed by dedicated, nuclear-encoded enzymes. Recent growing evidence suggests that mutations in these nuclear genes (nDNA), leading to incorrect maturation of mt-tRNAs, are a cause of human mitochondrial disease. Furthermore, mtDNA mutations in mt-tRNA genes, which may also affect mt-tRNA function, processing, and modification, are also frequently associated with human disease. In theory, all pathogenic mt-tRNA variants should be expected to affect only a single process, which is mitochondrial translation, albeit to various extents. However, the clinical manifestations of mitochondrial disorders linked to mutations in mt-tRNAs are extremely heterogeneous, ranging from defects of a single tissue to complex multisystem disorders. This review focuses on the current knowledge of nDNA coding for proteins involved in mt-tRNA maturation that have been linked to human mitochondrial pathologies. We further discuss the possibility that tissue specific regulation of mt-tRNA modifying enzymes could play an important role in the clinical heterogeneity observed for mitochondrial diseases caused by mutations in mt-tRNA genes.
Keywords: RNA modification; RNA processing; mitochondria; mitochondrial disease; post-transcriptional regulation; tRNA.
Figures

References
-
- Baruffini E., Dallabona C., Invernizzi F., Yarham J. W., Melchionda L., Blakely E. L., et al. (2013). MTO1 mutations are associated with hypertrophic cardiomyopathy and lactic acidosis and cause respiratory chain deficiency in humans and yeast. Hum. Mutat. 34 1501–1509 10.1002/humu.22393 - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

