Multiple genome segments determine virulence of bluetongue virus serotype 8
- PMID: 25822026
- PMCID: PMC4442542
- DOI: 10.1128/JVI.00395-15
Multiple genome segments determine virulence of bluetongue virus serotype 8
Abstract
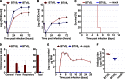
Bluetongue virus (BTV) causes bluetongue, a major hemorrhagic disease of ruminants. In order to investigate the molecular determinants of BTV virulence, we used a BTV8 strain minimally passaged in tissue culture (termed BTV8L in this study) and a derivative strain passaged extensively in tissue culture (BTV8H) in in vitro and in vivo studies. BTV8L was pathogenic in both IFNAR(-/-) mice and in sheep, while BTV8H was attenuated in both species. To identify genetic changes which led to BTV8H attenuation, we generated 34 reassortants between BTV8L and BTV8H. We found that partial attenuation of BTV8L in IFNAR(-/-) mice was achieved by simply replacing genomic segment 2 (Seg2, encoding VP2) or Seg10 (encoding NS3) with the BTV8H homologous segments. Fully attenuated viruses required at least two genome segments from BTV8H, including Seg2 with either Seg1 (encoding VP1), Seg6 (encoding VP6 and NS4), or Seg10 (encoding NS3). Conversely, full reversion of virulence of BTV8H required at least five genomic segments of BTV8L. We also demonstrated that BTV8H acquired an increased affinity for glycosaminoglycan receptors during passaging in cell culture due to mutations in its VP2 protein. Replication of BTV8H was relatively poor in interferon (IFN)-competent primary ovine endothelial cells compared to replication of BTV8L, and this phenotype was determined by several viral genomic segments, including Seg4 and Seg9. This study demonstrated that multiple viral proteins contribute to BTV8 virulence. VP2 and NS3 are primary determinants of BTV pathogenesis, but VP1, VP5, VP4, VP6, and VP7 also contribute to virulence.
Importance: Bluetongue is one of the major infectious diseases of ruminants, and it is listed as a notifiable disease by the World Organization for Animal Health (OIE). The clinical outcome of BTV infection varies considerably and depends on environmental and host- and virus-specific factors. Over the years, BTV serotypes/strains with various degrees of virulence (including nonpathogenic strains) have been described in different geographical locations. However, no data are available to correlate the BTV genotype to virulence. This study shows that BTV virulence is determined by different viral genomic segments. The data obtained will help to characterize thoroughly the pathogenesis of bluetongue. The possibility to determine the pathogenicity of virus isolates on the basis of their genome sequences will help in the design of control strategies that fit the risk posed by new emerging BTV strains.
Copyright © 2015, Janowicz et al.
Figures




References
-
- Price DA, Hardy WT. 1954. Isolation of the bluetongue virus from Texas sheep-Culicoides shown to be a vector. J Am Vet Med Assoc 124:255–258. - PubMed
-
- Mellor PS, Baylis M, Mertens PP. 2009. Bluetongue. Academic Press, London, United Kingdom.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

