Nrg1 is an injury-induced cardiomyocyte mitogen for the endogenous heart regeneration program in zebrafish
- PMID: 25830562
- PMCID: PMC4379493
- DOI: 10.7554/eLife.05871
Nrg1 is an injury-induced cardiomyocyte mitogen for the endogenous heart regeneration program in zebrafish
Abstract
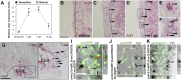
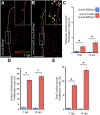
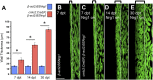
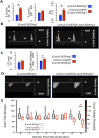
Heart regeneration is limited in adult mammals but occurs naturally in adult zebrafish through the activation of cardiomyocyte division. Several components of the cardiac injury microenvironment have been identified, yet no factor on its own is known to stimulate overt myocardial hyperplasia in a mature, uninjured animal. In this study, we find evidence that Neuregulin1 (Nrg1), previously shown to have mitogenic effects on mammalian cardiomyocytes, is sharply induced in perivascular cells after injury to the adult zebrafish heart. Inhibition of Erbb2, an Nrg1 co-receptor, disrupts cardiomyocyte proliferation in response to injury, whereas myocardial Nrg1 overexpression enhances this proliferation. In uninjured zebrafish, the reactivation of Nrg1 expression induces cardiomyocyte dedifferentiation, overt muscle hyperplasia, epicardial activation, increased vascularization, and causes cardiomegaly through persistent addition of wall myocardium. Our findings identify Nrg1 as a potent, induced mitogen for the endogenous adult heart regeneration program.
Keywords: Neuregulin1; cardiac hyperplasia; cardiomyocyte; developmental biology; epicardium; heart regeneration; stem cells; zebrafish.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures



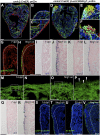
References
-
- Acharya A, Baek ST, Huang G, Eskiocak B, Goetsch S, Sung CY, Banfi S, Sauer MF, Olsen GS, Duffield JS, Olson EN, Tallquist MD. The bHLH transcription factor Tcf21 is required for lineage-specific EMT of cardiac fibroblast progenitors. Development. 2012;139:2139–2149. doi: 10.1242/dev.079970. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

