Ptch1 and Gli regulate Shh signalling dynamics via multiple mechanisms
- PMID: 25833741
- PMCID: PMC4396374
- DOI: 10.1038/ncomms7709
Ptch1 and Gli regulate Shh signalling dynamics via multiple mechanisms
Abstract
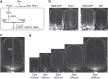
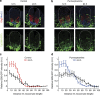
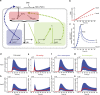
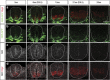
In the vertebrate neural tube, the morphogen Sonic Hedgehog (Shh) establishes a characteristic pattern of gene expression. Here we quantify the Shh gradient in the developing mouse neural tube and show that while the amplitude of the gradient increases over time, the activity of the pathway transcriptional effectors, Gli proteins, initially increases but later decreases. Computational analysis of the pathway suggests three mechanisms that could contribute to this adaptation: transcriptional upregulation of the inhibitory receptor Ptch1, transcriptional downregulation of Gli and the differential stability of active and inactive Gli isoforms. Consistent with this, Gli2 protein expression is downregulated during neural tube patterning and adaptation continues when the pathway is stimulated downstream of Ptch1. Moreover, the Shh-induced upregulation of Gli2 transcription prevents Gli activity levels from adapting in a different cell type, NIH3T3 fibroblasts, despite the upregulation of Ptch1. Multiple mechanisms therefore contribute to the intracellular dynamics of Shh signalling, resulting in different signalling dynamics in different cell types.
Figures


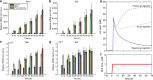
References
-
- Towers M., Wolpert L. & Tickle C. Gradients of signalling in the developing limb. Curr. Opin. Cell Biol. 24, 181–187 (2012) . - PubMed
-
- Dessaud E., McMahon A. P. & Briscoe J. Pattern formation in the vertebrate neural tube: a sonic hedgehog morphogen-regulated transcriptional network. Development 135, 2489–2503 (2008) . - PubMed
-
- Hui C.-C. & Angers S. Gli proteins in development and disease. Annu. Rev. Cell Dev. Biol. 27, 513–537 (2011) . - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

